MicroRNA gene dosage alterations and drug response in lung cancer
- PMID: 21541180
- PMCID: PMC3085440
- DOI: 10.1155/2011/474632
MicroRNA gene dosage alterations and drug response in lung cancer
Abstract
Chemotherapy resistance is a key contributor to the dismal prognoses for lung cancer patients. While the majority of studies have focused on sequence mutations and expression changes in protein-coding genes, recent reports have suggested that microRNA (miRNA) expression changes also play an influential role in chemotherapy response. However, the role of genetic alterations at miRNA loci in the context of chemotherapy response has yet to be investigated. In this study, we demonstrate the application of an integrative, multidimensional approach in order to identify miRNAs that are associated with chemotherapeutic resistance and sensitivity utilizing publicly available drug response, miRNA loci copy number, miRNA expression, and mRNA expression data from independent resources. By instigating a logical stepwise strategy, we have identified specific miRNAs that are associated with resistance to several chemotherapeutic agents and provide a proof of principle demonstration of how these various databases may be exploited to derive relevant pharmacogenomic results.
Figures



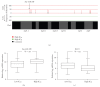
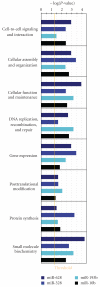
References
-
- Jemal A, Siegel R, Xu J, Ward E. Cancer statistics, 2010. CA: Cancer Journal for Clinicians. 2010;60(5):277–300. - PubMed
-
- Wakelee H, Belani CP. Optimizing first-line treatment options for patients with advanced NSCLC. Oncologist. 2005;10(3):1–10. - PubMed
-
- Scagliotti G, Hanna N, Fossella F, et al. The differential efficacy of pemetrexed according to NSCLC histology: a review of two phase III studies. Oncologist. 2009;14(3):253–263. - PubMed
-
- Fukuoka M, Yano S, Giaccone G, et al. Multi-institutional randomized phase II trial of gefitinib for previously treated patients with advanced non-small-cell lung cancer. Journal of Clinical Oncology. 2003;21(12):2237–2246. - PubMed
-
- Mok TS, Wu YL, Thongprasert S, et al. Gefitinib or carboplatin-paclitaxel in pulmonary adenocarcinoma. The New England Journal of Medicine. 2009;361(10):947–957. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

