Advanced mass spectrometry-based methods for the analysis of conformational integrity of biopharmaceutical products
- PMID: 21542797
- PMCID: PMC3375681
- DOI: 10.2174/138920111798357311
Advanced mass spectrometry-based methods for the analysis of conformational integrity of biopharmaceutical products
Abstract
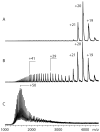
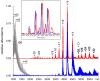
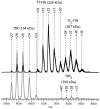
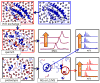
Mass spectrometry has already become an indispensable tool in the analytical armamentarium of the biopharmaceutical industry, although its current uses are limited to characterization of covalent structure of recombinant protein drugs. However, the scope of applications of mass spectrometry-based methods is beginning to expand to include characterization of the higher order structure and dynamics of biopharmaceutical products, a development which is catalyzed by the recent progress in mass spectrometry-based methods to study higher order protein structure. The two particularly promising methods that are likely to have the most significant and lasting impact in many areas of biopharmaceutical analysis, direct ESI MS and hydrogen/deuterium exchange, are focus of this article.
Figures
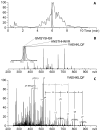
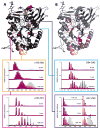
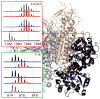
References
-
- Walsh G. Biopharmaceutical benchmarks 2010. Nature Biotechnol. 2010;28:917–294. - PubMed
-
- Lundblad RL. Approaches to the Conformational Analysis of Biopharmaceuticals. In: Lundblad RL, editor. Approaches to the conformational analysis of biopharmaceuticals. Boca Raton: CRC Press/Taylor & Francis; 2010.
-
- Kaltashov IA, Eyles SJ. Studies of biomolecular conformations and conformational dynamics by mass spectrometry. Mass Spectrom Rev. 2002;21:37–71. - PubMed
-
- Zhang Z, Pan H, Chen X. Mass spectrometry for structural characterization of therapeutic antibodies. Mass Spectrom Rev. 2009;28:147–176. - PubMed
-
- Srebalus-Barnes CA, Lim A. Applications of mass spectrometry for the structural characterization of recombinant protein pharmaceuticals. Mass Spectrom Rev. 2007;26:370–388. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

