Exploring the diversity of plant DNA viruses and their satellites using vector-enabled metagenomics on whiteflies
- PMID: 21544196
- PMCID: PMC3081322
- DOI: 10.1371/journal.pone.0019050
Exploring the diversity of plant DNA viruses and their satellites using vector-enabled metagenomics on whiteflies
Abstract
Current knowledge of plant virus diversity is biased towards agents of visible and economically important diseases. Less is known about viruses that have not caused major diseases in crops, or viruses from native vegetation, which are a reservoir of biodiversity that can contribute to viral emergence. Discovery of these plant viruses is hindered by the traditional approach of sampling individual symptomatic plants. Since many damaging plant viruses are transmitted by insect vectors, we have developed "vector-enabled metagenomics" (VEM) to investigate the diversity of plant viruses. VEM involves sampling of insect vectors (in this case, whiteflies) from plants, followed by purification of viral particles and metagenomic sequencing. The VEM approach exploits the natural ability of highly mobile adult whiteflies to integrate viruses from many plants over time and space, and leverages the capability of metagenomics for discovering novel viruses. This study utilized VEM to describe the DNA viral community from whiteflies (Bemisia tabaci) collected from two important agricultural regions in Florida, USA. VEM successfully characterized the active and abundant viruses that produce disease symptoms in crops, as well as the less abundant viruses infecting adjacent native vegetation. PCR assays designed from the metagenomic sequences enabled the complete sequencing of four novel begomovirus genome components, as well as the first discovery of plant virus satellites in North America. One of the novel begomoviruses was subsequently identified in symptomatic Chenopodium ambrosiodes from the same field site, validating VEM as an effective method for proactive monitoring of plant viruses without a priori knowledge of the pathogens. This study demonstrates the power of VEM for describing the circulating viral community in a given region, which will enhance our understanding of plant viral diversity, and facilitate emerging plant virus surveillance and management of viral diseases.
Conflict of interest statement
Figures

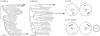
References
-
- Nawaz-Ul-Rehman MS, Fauquet CM. Evolution of geminiviruses and their satellites. FEBS Letters. 2009;583:1825–1832. - PubMed
-
- Zhou X, Liu Y, Calvert L, Munoz C, Otim-Nape GW, et al. Evidence that DNA-A of a geminivirus associated with severe cassava mosaic disease in Uganda has arisen by interspecific recombination. Journal of General Virology. 1997;78:2101–2111. - PubMed
-
- Zhou X, Liu Y, Robinson DJ, Harrison BD. Four DNA-A variants among Pakistani isolates of cotton leaf curl virus and their affinities to DNA-A of geminivirus isolates from okra. Journal of General Virology. 1998;79:915–923. - PubMed
-
- Brown JK. Molecular markers for the identification and global tracking of whitefly vector-Begomovirus complexes. Virus Research. 2000;71:233–260. - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources

