Functional selectivity of adenosine receptor ligands
- PMID: 21544511
- PMCID: PMC3146648
- DOI: 10.1007/s11302-011-9232-0
Functional selectivity of adenosine receptor ligands
Abstract
Adenosine receptors are plasma membrane proteins that transduce an extracellular signal into the interior of the cell. Basically every mammalian cell expresses at least one of the four adenosine receptor subtypes. Recent insight in signal transduction cascades teaches us that the current classification of receptor ligands into agonists, antagonists, and inverse agonists relies very much on the experimental setup that was used. Upon activation of the receptors by the ubiquitous endogenous ligand adenosine they engage classical G protein-mediated pathways, resulting in production of second messengers and activation of kinases. Besides this well-described G protein-mediated signaling pathway, adenosine receptors activate scaffold proteins such as β-arrestins. Using innovative and sensitive experimental tools, it has been possible to detect ligands that preferentially stimulate the β-arrestin pathway over the G protein-mediated signal transduction route, or vice versa. This phenomenon is referred to as functional selectivity or biased signaling and implies that an antagonist for one pathway may be a full agonist for the other signaling route. Functional selectivity makes it necessary to redefine the functional properties of currently used adenosine receptor ligands and opens possibilities for new and more selective ligands. This review focuses on the current knowledge of functionally selective adenosine receptor ligands and on G protein-independent signaling of adenosine receptors through scaffold proteins.
Figures
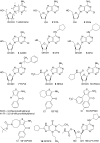


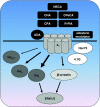
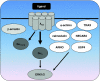
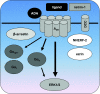
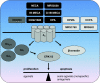
References
-
- Kristiansen K. Molecular mechanisms of ligand binding, signaling, and regulation within the superfamily of G-protein-coupled receptors: molecular modeling and mutagenesis approaches to receptor structure and function. Pharmacol Ther. 2004;103(1):21–80. - PubMed
-
- Oakley RH, Laporte SA, Holt JA, Caron MG, Barak LS. Differential affinities of visual arrestin, βarrestin1, and βarrestin2 for G protein-coupled receptors delineate two major classes of receptors. J Biol Chem. 2000;275(22):17201–17210. - PubMed
LinkOut - more resources
Full Text Sources
Other Literature Sources

