Recombinant expression and functional analysis of proteases from Streptococcus pneumoniae, Bacillus anthracis, and Yersinia pestis
- PMID: 21545736
- PMCID: PMC3113736
- DOI: 10.1186/1471-2091-12-17
Recombinant expression and functional analysis of proteases from Streptococcus pneumoniae, Bacillus anthracis, and Yersinia pestis
Abstract
Background: Uncharacterized proteases naturally expressed by bacterial pathogens represents important topic in infectious disease research, because these enzymes may have critical roles in pathogenicity and cell physiology. It has been observed that cloning, expression and purification of proteases often fail due to their catalytic functions which, in turn, cause toxicity in the E. coli heterologous host.
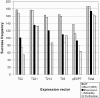
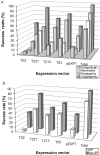
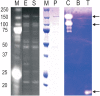
Results: In order to address this problem systematically, a modified pipeline of our high-throughput protein expression and purification platform was developed. This included the use of a specific E. coli strain, BL21(DE3) pLysS to tightly control the expression of recombinant proteins and various expression vectors encoding fusion proteins to enhance recombinant protein solubility. Proteases fused to large fusion protein domains, maltosebinding protein (MBP), SP-MBP which contains signal peptide at the N-terminus of MBP, disulfide oxidoreductase (DsbA) and Glutathione S-transferase (GST) improved expression and solubility of proteases. Overall, 86.1% of selected protease genes including hypothetical proteins were expressed and purified using a combination of five different expression vectors. To detect novel proteolytic activities, zymography and fluorescence-based assays were performed and the protease activities of more than 46% of purified proteases and 40% of hypothetical proteins that were predicted to be proteases were confirmed.
Conclusions: Multiple expression vectors, employing distinct fusion tags in a high throughput pipeline increased overall success rates in expression, solubility and purification of proteases. The combinatorial functional analysis of the purified proteases using fluorescence assays and zymography confirmed their function.
Figures


References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

