Minireview: The roles of small RNA pathways in reproductive medicine
- PMID: 21546411
- PMCID: PMC3146245
- DOI: 10.1210/me.2011-0099
Minireview: The roles of small RNA pathways in reproductive medicine
Abstract
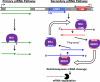
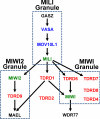
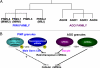
The discovery of small noncoding RNA, including P-element-induced wimpy testis-interacting RNA, small interfering RNA, and microRNA, has energized research in reproductive medicine. In the two decades since the identification of small RNA, first in Caenorhabditis elegans and then in other animals, scientists in many disciplines have made significant progress in elucidating their biology. A powerful battery of tools, including knockout mice and small RNA mimics and antagonists, has facilitated investigation into the functional roles and therapeutic potential of these small RNA pathways. Current data indicate that small RNA play significant roles in normal development and physiology and pathological conditions of the reproductive tracts of females and males. Biologically plausible mRNA targets for these microRNA are aggressively being discovered. The next phase of research will focus on elucidating the clinical utility of small RNA-selective agonists and antagonists.
Figures

Similar articles
-
Biochemical principles of small RNA pathways.Annu Rev Biochem. 2010;79:295-319. doi: 10.1146/annurev.biochem.052208.151733. Annu Rev Biochem. 2010. PMID: 20205586 Review.
-
The benefits of basic research: advances in reproductive physiology.Popul Briefs. 1995 Jun;1(2):1-2. Popul Briefs. 1995. PMID: 12319542
-
MicroRNA: a small molecule with a big biological impact.Microrna. 2012;1(1):1. doi: 10.2174/2211536611201010001. Microrna. 2012. PMID: 25048083
-
Small RNAs in mammalian germline: Tiny for immortal.Differentiation. 2010 Mar;79(3):141-6. doi: 10.1016/j.diff.2009.11.002. Differentiation. 2010. PMID: 20227007 Review.
-
Small RNAs in germline development.Curr Top Dev Biol. 2013;102:159-205. doi: 10.1016/B978-0-12-416024-8.00006-4. Curr Top Dev Biol. 2013. PMID: 23287033 Review.
Cited by
-
MicroRNAs--mediators of myometrial contractility during pregnancy and labour.Nat Rev Endocrinol. 2013 Jul;9(7):391-401. doi: 10.1038/nrendo.2013.96. Epub 2013 May 14. Nat Rev Endocrinol. 2013. PMID: 23669656 Review.
-
Structural basis for arginine methylation-independent recognition of PIWIL1 by TDRD2.Proc Natl Acad Sci U S A. 2017 Nov 21;114(47):12483-12488. doi: 10.1073/pnas.1711486114. Epub 2017 Nov 8. Proc Natl Acad Sci U S A. 2017. PMID: 29118143 Free PMC article.
-
Minireview: miRomics in endocrinology: a novel approach for modeling endocrine diseases.Mol Endocrinol. 2013 Apr;27(4):573-85. doi: 10.1210/me.2012-1220. Epub 2013 Jan 24. Mol Endocrinol. 2013. PMID: 23349525 Free PMC article. Review.
-
The microRNA (miR)-199a/214 cluster mediates opposing effects of progesterone and estrogen on uterine contractility during pregnancy and labor.Mol Endocrinol. 2012 Nov;26(11):1857-67. doi: 10.1210/me.2012-1199. Epub 2012 Sep 12. Mol Endocrinol. 2012. PMID: 22973051 Free PMC article.
-
Identifying functional cancer-specific miRNA-mRNA interactions in testicular germ cell tumor.J Theor Biol. 2016 Sep 7;404:82-96. doi: 10.1016/j.jtbi.2016.05.026. Epub 2016 May 25. J Theor Biol. 2016. PMID: 27235586 Free PMC article.
References
-
- Fire A, Xu S, Montgomery MK, Kostas SA, Driver SE, Mello CC. 1998. Potent and specific genetic interference by double-stranded RNA in Caenorhabditis elegans. Nature 391:806–811 - PubMed
-
- Girard A, Sachidanandam R, Hannon GJ, Carmell MA. 2006. A germline-specific class of small RNAs binds mammalian Piwi proteins. Nature 442:199–202 - PubMed
-
- Lau NC, Seto AG, Kim J, Kuramochi-Miyagawa S, Nakano T, Bartel DP, Kingston RE. 2006. Characterization of the piRNA complex from rat testes. Science 313:363–367 - PubMed
-
- Aravin A, Gaidatzis D, Pfeffer S, Lagos-Quintana M, Landgraf P, Iovino N, Morris P, Brownstein MJ, Kuramochi-Miyagawa S, Nakano T, Chien M, Russo JJ, Ju J, Sheridan R, Sander C, Zavolan M, Tuschl T. 2006. A novel class of small RNAs bind to MILI protein in mouse testes. Nature 442:203–207 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
- 5K12HD050128/HD/NICHD NIH HHS/United States
- 5T32HD007165/HD/NICHD NIH HHS/United States
- R03 HD077495/HD/NICHD NIH HHS/United States
- U01HD060496/HD/NICHD NIH HHS/United States
- R01 CA060651/CA/NCI NIH HHS/United States
- R01CA060651/CA/NCI NIH HHS/United States
- K12 HD050128/HD/NICHD NIH HHS/United States
- R01HD033438/HD/NICHD NIH HHS/United States
- R01 HD032067/HD/NICHD NIH HHS/United States
- T32 HD007165/HD/NICHD NIH HHS/United States
- R01 HD033438/HD/NICHD NIH HHS/United States
- U01 HD060496/HD/NICHD NIH HHS/United States
- U54HD0077495/HD/NICHD NIH HHS/United States

