Functional context, biosynthesis, and genetic encoding of pyrrolysine
- PMID: 21550296
- PMCID: PMC3119745
- DOI: 10.1016/j.mib.2011.04.001
Functional context, biosynthesis, and genetic encoding of pyrrolysine
Abstract
In Methanosarcina spp., amber codons in methylamine methyltransferase genes are translated as the 22nd amino acid, pyrrolysine. The responsible pyl genes plus amber-codon containing methyltransferase genes have been identified in four archaeal and five bacterial genera, including one human pathogen. In Escherichia coli, the recombinant pylBCD gene products biosynthesize pyrrolysine from two molecules of lysine and the pylTS gene products direct pyrrolysine incorporation into protein. In the proposed biosynthetic pathway, PylB forms methylornithine from lysine, which is joined to another lysine by PylC, and oxidized to pyrrolysine by PylD. Structures of the catalytic domain of pyrrolysyl-tRNA synthetase (archaeal PylS or bacterial PylSc) revealed binding sites for tRNAPyl and pyrrolysine. PylS and tRNAPyl are now being exploited as an orthogonal pair in recombinant systems for introduction of useful modified amino acids into proteins.
Copyright © 2011 Elsevier Ltd. All rights reserved.
Figures


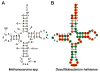
References
-
- Shima S, Warkentin E, Thauer RK, Ermler U. Structure and function of enzymes involved in the methanogenic pathway utilizing carbon dioxide and molecular hydrogen. J Biosci Bioeng. 2002;93:519–530. - PubMed
-
- Srinivasan G, James CM, Krzycki JA. Pyrrolysine encoded by UAG in Archaea: charging of a UAG-decoding specialized tRNA. Science. 2002;296:1459–1462. - PubMed
-
- Hao B, Gong W, Ferguson TK, James CM, Krzycki JA, Chan MK. A new UAG-encoded residue in the structure of a methanogen methyltransferase. Science. 2002;296:1462–1466. - PubMed
-
- Ferguson DJ, Jr, Gorlatova N, Grahame DA, Krzycki JA. Reconstitution of dimethylamine:coenzyme M methyl transfer with a discrete corrinoid protein and two methyltransferases purified from Methanosarcina barkeri. J Biol Chem. 2000;275:29053–29060. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

