mirConnX: condition-specific mRNA-microRNA network integrator
- PMID: 21558324
- PMCID: PMC3125733
- DOI: 10.1093/nar/gkr276
mirConnX: condition-specific mRNA-microRNA network integrator
Abstract
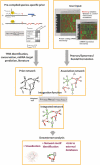
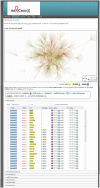
mirConnX is a user-friendly web interface for inferring, displaying and parsing mRNA and microRNA (miRNA) gene regulatory networks. mirConnX combines sequence information with gene expression data analysis to create a disease-specific, genome-wide regulatory network. A prior, static network has been constructed for all human and mouse genes. It consists of computationally predicted transcription factor (TF)-gene associations and miRNA target predictions. The prior network is supplemented with known interactions from the literature. Dynamic TF- and miRNA-gene associations are inferred from user-provided expression data using an association measure of choice. The static and dynamic networks are then combined using an integration function with user-specified weights. Visualization of the network and subsequent analysis are provided via a very responsive graphic user interface. Two organisms are currently supported: Homo sapiens and Mus musculus. The intuitive user interface and large database make mirConnX a useful tool for clinical scientists for hypothesis generation and explorations. mirConnX is freely available for academic use at http://www.benoslab.pitt.edu/mirconnx.
Figures
Similar articles
-
DIANA-mirExTra v2.0: Uncovering microRNAs and transcription factors with crucial roles in NGS expression data.Nucleic Acids Res. 2016 Jul 8;44(W1):W128-34. doi: 10.1093/nar/gkw455. Epub 2016 May 20. Nucleic Acids Res. 2016. PMID: 27207881 Free PMC article.
-
ComiR: Combinatorial microRNA target prediction tool.Nucleic Acids Res. 2013 Jul;41(Web Server issue):W159-64. doi: 10.1093/nar/gkt379. Epub 2013 May 22. Nucleic Acids Res. 2013. PMID: 23703208 Free PMC article.
-
MAGIA, a web-based tool for miRNA and Genes Integrated Analysis.Nucleic Acids Res. 2010 Jul;38(Web Server issue):W352-9. doi: 10.1093/nar/gkq423. Epub 2010 May 19. Nucleic Acids Res. 2010. PMID: 20484379 Free PMC article.
-
Participation of microRNAs in human interactome: extraction of microRNA-microRNA regulations.Mol Biosyst. 2011 Jun;7(6):1966-73. doi: 10.1039/c0mb00347f. Epub 2011 Apr 11. Mol Biosyst. 2011. PMID: 21483898
-
CircuitsDB: a database of mixed microRNA/transcription factor feed-forward regulatory circuits in human and mouse.BMC Bioinformatics. 2010 Aug 23;11:435. doi: 10.1186/1471-2105-11-435. BMC Bioinformatics. 2010. PMID: 20731828 Free PMC article.
Cited by
-
Tuning the engine: an introduction to resources on post-transcriptional regulation of gene expression.RNA Biol. 2012 Oct;9(10):1224-32. doi: 10.4161/rna.22035. Epub 2012 Sep 20. RNA Biol. 2012. PMID: 22995832 Free PMC article. Review.
-
Integrated miRNA and mRNA Analysis of Time Series Microarray Data.ACM BCB. 2014;2014:122-127. doi: 10.1145/2649387.2649411. ACM BCB. 2014. PMID: 25988189 Free PMC article.
-
An integrated approach to characterize transcription factor and microRNA regulatory networks involved in Schwann cell response to peripheral nerve injury.BMC Genomics. 2013 Feb 6;14:84. doi: 10.1186/1471-2164-14-84. BMC Genomics. 2013. PMID: 23387820 Free PMC article.
-
Integrated Genomics Reveals Convergent Transcriptomic Networks Underlying Chronic Obstructive Pulmonary Disease and Idiopathic Pulmonary Fibrosis.Am J Respir Crit Care Med. 2016 Oct 15;194(8):948-960. doi: 10.1164/rccm.201510-2026OC. Am J Respir Crit Care Med. 2016. PMID: 27104832 Free PMC article.
-
Impact of microRNA regulation on variation in human gene expression.Genome Res. 2012 Jul;22(7):1243-54. doi: 10.1101/gr.132514.111. Epub 2012 Mar 28. Genome Res. 2012. PMID: 22456605 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

