Multizone paper platform for 3D cell cultures
- PMID: 21573103
- PMCID: PMC3089608
- DOI: 10.1371/journal.pone.0018940
Multizone paper platform for 3D cell cultures
Abstract
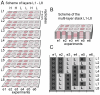
In vitro 3D culture is an important model for tissues in vivo. Cells in different locations of 3D tissues are physiologically different, because they are exposed to different concentrations of oxygen, nutrients, and signaling molecules, and to other environmental factors (temperature, mechanical stress, etc). The majority of high-throughput assays based on 3D cultures, however, can only detect the average behavior of cells in the whole 3D construct. Isolation of cells from specific regions of 3D cultures is possible, but relies on low-throughput techniques such as tissue sectioning and micromanipulation. Based on a procedure reported previously ("cells-in-gels-in-paper" or CiGiP), this paper describes a simple method for culture of arrays of thin planar sections of tissues, either alone or stacked to create more complex 3D tissue structures. This procedure starts with sheets of paper patterned with hydrophobic regions that form 96 hydrophilic zones. Serial spotting of cells suspended in extracellular matrix (ECM) gel onto the patterned paper creates an array of 200 micron-thick slabs of ECM gel (supported mechanically by cellulose fibers) containing cells. Stacking the sheets with zones aligned on top of one another assembles 96 3D multilayer constructs. De-stacking the layers of the 3D culture, by peeling apart the sheets of paper, "sections" all 96 cultures at once. It is, thus, simple to isolate 200-micron-thick cell-containing slabs from each 3D culture in the 96-zone array. Because the 3D cultures are assembled from multiple layers, the number of cells plated initially in each layer determines the spatial distribution of cells in the stacked 3D cultures. This capability made it possible to compare the growth of 3D tumor models of different spatial composition, and to examine the migration of cells in these structures.
Conflict of interest statement
Figures



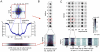



Similar articles
-
Polymer-based mesh as supports for multi-layered 3D cell culture and assays.Biomaterials. 2014 Jan;35(1):259-68. doi: 10.1016/j.biomaterials.2013.09.049. Epub 2013 Oct 2. Biomaterials. 2014. PMID: 24095253 Free PMC article.
-
Platform for high-throughput testing of the effect of soluble compounds on 3D cell cultures.Anal Chem. 2013 Sep 3;85(17):8085-94. doi: 10.1021/ac400161j. Epub 2013 Aug 16. Anal Chem. 2013. PMID: 23952342 Free PMC article.
-
Paper-supported 3D cell culture for tissue-based bioassays.Proc Natl Acad Sci U S A. 2009 Nov 3;106(44):18457-62. doi: 10.1073/pnas.0910666106. Epub 2009 Oct 21. Proc Natl Acad Sci U S A. 2009. PMID: 19846768 Free PMC article.
-
Cancer Cytokines and the Relevance of 3D Cultures for Studying Those Implicated in Human Cancers.J Cell Biochem. 2017 Sep;118(9):2544-2558. doi: 10.1002/jcb.25970. Epub 2017 May 3. J Cell Biochem. 2017. PMID: 28262975 Review.
-
Layers and Multilayers of Self-Assembled Polymers: Tunable Engineered Extracellular Matrix Coatings for Neural Cell Growth.Langmuir. 2018 Jul 31;34(30):8709-8730. doi: 10.1021/acs.langmuir.7b04108. Epub 2018 Mar 12. Langmuir. 2018. PMID: 29481757 Review.
Cited by
-
Osteogenic differentiation of preosteoblasts on a hemostatic gelatin sponge.Sci Rep. 2016 Sep 12;6:32884. doi: 10.1038/srep32884. Sci Rep. 2016. PMID: 27616161 Free PMC article.
-
Flexible and disposable paper- and plastic-based gel micropads for nematode handling, imaging, and chemical testing.APL Bioeng. 2017 Oct 9;1(1):016102. doi: 10.1063/1.5005829. eCollection 2017 Dec. APL Bioeng. 2017. PMID: 31069282 Free PMC article.
-
Polymer-based mesh as supports for multi-layered 3D cell culture and assays.Biomaterials. 2014 Jan;35(1):259-68. doi: 10.1016/j.biomaterials.2013.09.049. Epub 2013 Oct 2. Biomaterials. 2014. PMID: 24095253 Free PMC article.
-
Microfluidics-based 3D cell culture models: Utility in novel drug discovery and delivery research.Bioeng Transl Med. 2016 Jul 5;1(1):63-81. doi: 10.1002/btm2.10013. eCollection 2016 Mar. Bioeng Transl Med. 2016. PMID: 29313007 Free PMC article. Review.
-
Current methods for studying metastatic potential of tumor cells.Cancer Cell Int. 2022 Dec 9;22(1):394. doi: 10.1186/s12935-022-02801-w. Cancer Cell Int. 2022. PMID: 36494720 Free PMC article. Review.
References
-
- Abbott A. Cell culture: Biology's new dimension. Nature. 2003;424:870–872. - PubMed
-
- Sutherland RM. Cell and Environment Interactions in Tumor Microregions-the Multicell Spheroid Model. Science. 1988;240:177–184. - PubMed
-
- Yamada KM, Cukierman E. Modeling tissue morphogenesis and cancer in 3D. Cell. 2007;130:601–610. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

