Peptide-protein interactions suggest that acetylation of lysines 381 and 382 of p53 is important for positive coactivator 4-p53 interaction
- PMID: 21586571
- PMCID: PMC3137081
- DOI: 10.1074/jbc.M110.205328
Peptide-protein interactions suggest that acetylation of lysines 381 and 382 of p53 is important for positive coactivator 4-p53 interaction
Abstract
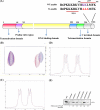
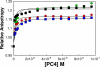
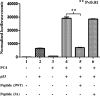
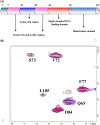
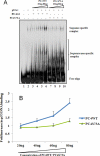
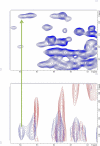
The human transcriptional positive coactivator 4 (PC4) activates several p53-dependent genes. It has been demonstrated that this is a consequence of direct interaction with p53. Previously, we have concluded that PC4 interacts mainly with the C-terminal negative regulatory domain of p53 through its DNA binding C-terminal half. NMR chemical shift perturbation studies with peptide fragments indicated that amino acids 380-386 of p53 are crucial for interaction with PC4. This was verified by fluorescence anisotropy and sedimentation velocity studies. A peptide consisting of p53-(380-386) sequence, when attached to a cell penetration tag and nuclear localization signal, localizes to the nucleus and inhibits luciferase gene expression from a transfected plasmid carrying a Luc gene under a p53-dependent promoter. Acetylation of lysine 382/381 enhanced the binding of this peptide to PC4 by about an order of magnitude. NMR and mutagenesis studies indicated that serine 73 of PC4 is an important residue for recognition of p53. Intermolecular nuclear Overhauser effect placed aspartate 76 in the vicinity of lysine 381, indicating that the region around residues 73-76 of PC4 is important for p53 recognition. We conclude that the 380-386 region of p53 interacts with the region around residues 73-76 of PC4, and acetylation of lysine 382/381 of p53 may play an important role in modulating p53-PC4 interaction and as a consequence PC4 mediated activation of p53 target genes.
Figures



References
-
- Momand J., Zambetti G. P., Olson D. C., George D., Levine A. J. (1992) Cell 69, 1237–1245 - PubMed
-
- Dornan D., Wertz I., Shimizu H., Arnott D., Frantz G. D., Dowd P., O'Rourke K., Koeppen H., Dixit V. M. (2004) Nature 429, 86–92 - PubMed
-
- Leng R. P., Lin Y., Ma W., Wu H., Lemmers B., Chung S., Parant J. M., Lozano G., Hakem R., Benchimol S. (2003) Cell 112, 779–791 - PubMed
-
- Haupt Y., Maya R., Kazaz A., Oren M. (1997) Nature 387, 296–299 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

