BAC library resources for map-based cloning and physical map construction in barley (Hordeum vulgare L.)
- PMID: 21595870
- PMCID: PMC3224359
- DOI: 10.1186/1471-2164-12-247
BAC library resources for map-based cloning and physical map construction in barley (Hordeum vulgare L.)
Abstract
Background: Although second generation sequencing (2GS) technologies allow re-sequencing of previously gold-standard-sequenced genomes, whole genome shotgun sequencing and de novo assembly of large and complex eukaryotic genomes is still difficult. Availability of a genome-wide physical map is therefore still a prerequisite for whole genome sequencing for genomes like barley. To start such an endeavor, large insert genomic libraries, i.e. Bacterial Artificial Chromosome (BAC) libraries, which are unbiased and representing deep haploid genome coverage, need to be ready in place.
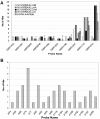
Result: Five new BAC libraries were constructed for barley (Hordeum vulgare L.) cultivar Morex. These libraries were constructed in different cloning sites (HindIII, EcoRI, MboI and BstXI) of the respective vectors. In order to enhance unbiased genome representation and to minimize the number of gaps between BAC contigs, which are often due to uneven distribution of restriction sites, a mechanically sheared library was also generated. The new BAC libraries were fully characterized in depth by scrutinizing the major quality parameters such as average insert size, degree of contamination (plate wide, neighboring, and chloroplast), empty wells and off-scale clones (clones with <30 or >250 fragments). Additionally a set of gene-based probes were hybridized to high density BAC filters and showed that genome coverage of each library is between 2.4 and 6.6 X.
Conclusion: BAC libraries representing >20 haploid genomes are available as a new resource to the barley research community. Systematic utilization of these libraries in high-throughput BAC fingerprinting should allow developing a genome-wide physical map for the barley genome, which will be instrumental for map-based gene isolation and genome sequencing.
Figures



References
-
- Peterson DG, Tomkins JP, Frisch DA, Wing RA, Paterson AH. Construction of plant bacterial artificial chromosomes (BAC) libraries: an illustrated guide. Journal of Agricultural genomics. 2000;5
-
- Vilarinhos AD, Piffanelli P, Lagoda P, Thibivilliers S, Sabau X, Carreel F, D'Hont A. Construction and characterization of a bacterial artificial chromosome library of banana (Musa acuminata Colla) Theor Appl Genet. 2003;106(6):1102–1106. - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources

