Compensatory signals associated with the activation of human GC 5' splice sites
- PMID: 21609956
- PMCID: PMC3167603
- DOI: 10.1093/nar/gkr306
Compensatory signals associated with the activation of human GC 5' splice sites
Abstract
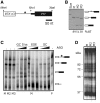
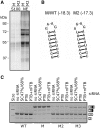
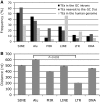
GC 5' splice sites (5'ss) are present in ∼1% of human introns, but factors promoting their efficient selection are poorly understood. Here, we describe a case of X-linked agammaglobulinemia resulting from a GC 5'ss activated by a mutation in BTK intron 3. This GC 5'ss was intrinsically weak, yet it was selected in >90% primary transcripts in the presence of a strong and intact natural GT counterpart. We show that efficient selection of this GC 5'ss required a high density of GAA/CAA-containing splicing enhancers in the exonized segment and was promoted by SR proteins 9G8, Tra2β and SC35. The GC 5'ss was efficiently inhibited by splice-switching oligonucleotides targeting either the GC 5'ss itself or the enhancer. Comprehensive analysis of natural GC-AG introns and previously reported pathogenic GC 5'ss showed that their efficient activation was facilitated by higher densities of splicing enhancers and lower densities of silencers than their GT 5'ss equivalents. Removal of the GC-AG introns was promoted to a minor extent by the splice-site strength of adjacent exons and inhibited by flanking Alu repeats, with the first downstream Alus located on average at a longer distance from the GC 5'ss than other transposable elements. These results provide new insights into the splicing code that governs selection of noncanonical splice sites.
Figures


References
-
- Wahl MC, Will CL, Lührmann R. The spliceosome: design principles of a dynamic RNP machine. Cell. 2009;136:701–718. - PubMed
-
- Burge CB, Tuschl T, Sharp PA. In: The RNA World. Gesteland RF, Cech TR, Atkins JF, editors. New York: Cold Spring Harbor Laboratory Press; 1999. pp. 525–560.
Publication types
MeSH terms
Substances
Supplementary concepts
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

