Glia maturation factor-γ mediates neutrophil chemotaxis
- PMID: 21653232
- PMCID: PMC3157903
- DOI: 10.1189/jlb.0710424
Glia maturation factor-γ mediates neutrophil chemotaxis
Abstract
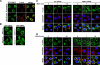
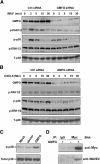
Chemotaxis is fundamental to the directional migration of neutrophils toward endogenous and exogenous chemoattractants. Recent studies have demonstrated that ADF/cofilin superfamily members play important roles in reorganizing the actin cytoskeleton by disassembling actin filaments. GMFG, a novel ADF/cofilin superfamily protein that is expressed in inflammatory cells, has been implicated in regulating actin reorganization in microendothelial cells, but its function in neutrophils remains unclear. Here, we show that GMFG is an important regulator for cell migration and polarity in neutrophils. Knockdown of endogenous GMFG impaired fMLF- and IL-8 (CXCL8)-induced chemotaxis in dHL-60 cells. GMFG knockdown attenuated the formation of lamellipodia at the leading edge of cells exposed to fMLF or CXCL8, as well as the phosphorylation of p38 and PAK1/2 in response to fMLF or CXCL8. Live cell imaging revealed that GMFG was recruited to the leading edge of cells in response to fMLF, as well as CXCL8. Overexpression of GMFG enhanced phosphorylation of p38 but not of PAK1/2 in dHL-60 cells. In addition, we found that GMFG is associated with WAVE2. Taken together, our findings suggest that GMFG is a novel factor in regulating neutrophil chemotaxis by modulating actin cytoskeleton reorganization.
Figures



References
-
- Nathan C. (2006) Neutrophils and immunity: challenges and opportunities. Nat. Rev. Immunol. 6, 173–182 - PubMed
-
- Ridley A. J., Schwartz M. A., Burridge K., Firtel R. A., Ginsberg M. H., Borisy G., Parsons J. T., Horwitz A. R. (2003) Cell migration: integrating signals from front to back. Science 302, 1704–1709 - PubMed
-
- Van Haastert P. J., Devreotes P. N. (2004) Chemotaxis: signaling the way forward. Nat. Rev. Mol. Cell Biol. 5, 626–634 - PubMed
-
- Devreotes P., Janetopoulos C. (2003) Eukaryotic chemotaxis: distinctions between directional sensing and polarization. J. Biol. Chem. 278, 20445–20448 - PubMed
-
- Webb D. J., Parsons J. T., Horwitz A. F. (2002) Adhesion assembly, disassembly and turnover in migrating cells—over and over and over again. Nat. Cell Biol. 4, E97–E100 - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

