De novo discovery of mutated driver pathways in cancer
- PMID: 21653252
- PMCID: PMC3266044
- DOI: 10.1101/gr.120477.111
De novo discovery of mutated driver pathways in cancer
Abstract
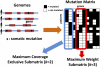
Next-generation DNA sequencing technologies are enabling genome-wide measurements of somatic mutations in large numbers of cancer patients. A major challenge in the interpretation of these data is to distinguish functional "driver mutations" important for cancer development from random "passenger mutations." A common approach for identifying driver mutations is to find genes that are mutated at significant frequency in a large cohort of cancer genomes. This approach is confounded by the observation that driver mutations target multiple cellular signaling and regulatory pathways. Thus, each cancer patient may exhibit a different combination of mutations that are sufficient to perturb these pathways. This mutational heterogeneity presents a problem for predicting driver mutations solely from their frequency of occurrence. We introduce two combinatorial properties, coverage and exclusivity, that distinguish driver pathways, or groups of genes containing driver mutations, from groups of genes with passenger mutations. We derive two algorithms, called Dendrix, to find driver pathways de novo from somatic mutation data. We apply Dendrix to analyze somatic mutation data from 623 genes in 188 lung adenocarcinoma patients, 601 genes in 84 glioblastoma patients, and 238 known mutations in 1000 patients with various cancers. In all data sets, we find groups of genes that are mutated in large subsets of patients and whose mutations are approximately exclusive. Our Dendrix algorithms scale to whole-genome analysis of thousands of patients and thus will prove useful for larger data sets to come from The Cancer Genome Atlas (TCGA) and other large-scale cancer genome sequencing projects.
Figures

References
-
- Backlund LM, Nilsson BR, Goike HM, Schmidt EE, Liu L, Ichimura K, Collins VP 2003. Short postoperative survival for glioblastoma patients with a dysfunctional Rb1 pathway in combination with no wild-type PTEN. Clin Cancer Res 9: 4151–4158 - PubMed
-
- Ben-Dor A, Chor B, Karp RM, Yakhini Z 2003. Discovering local structure in gene expression data: The order-preserving submatrix problem. J Comput Biol 10: 373–384 - PubMed
-
- Bradley JR, Farnsworth DL 2009. Testing for mutual exclusivity. J Appl Stat 36: 1307–1314
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources
