N-methylation of the amide bond by methyltransferase asm10 in ansamitocin biosynthesis
- PMID: 21681880
- PMCID: PMC3391011
- DOI: 10.1002/cbic.201100062
N-methylation of the amide bond by methyltransferase asm10 in ansamitocin biosynthesis
Abstract
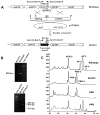
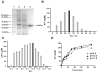
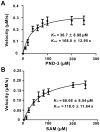
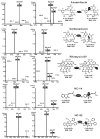
Ansamitocins are potent antitumor agents produced by Actinosynnema pretiosum. As deduced from their structures, an N-methylation on the amide bond is required among the various modifications. The protein encoded by asm10 belongs to the SAM-dependent methyltransferase family. Through gene inactivation and complementation, asm10 was proved to be responsible for the N-methylation of ansamitocins. Asm10 is a 33.0 kDa monomer, as determined by gel filtration. By using N-desmethyl-ansamitocin P-3 as substrate, the optimal temperature and pH for Asm10 catalysis were determined to be 32 °C and 10.0, respectively. Asm10 also showed broad substrate flexibility toward other N-desmethyl-ansamycins and synthetic indolin-2-ones. Through site-directed mutagenesis, Asp154 and Leu155 of Asm10 were confirmed to be essential for its catalysis, possibly through the binding of SAM. The characterization of this unique N-methyltransferase has enriched the toolbox for engineering N-methylated derivatives from both natural and synthetic compounds; this will allow known potential drugs to be modified.
Copyright © 2011 WILEY-VCH Verlag GmbH & Co. KGaA, Weinheim.
Figures

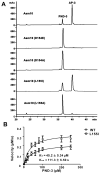
References
-
- Higashide E, Asai M, Ootsu K, Tanida S, Kozai Y, Hasegawa T, Kishi T, Sugino Y, Yoneda M. Nature. 1977;270:721–722. - PubMed
-
- Tanida S, Hasegawa T, Hatano K, Higashide E, Yoneda M. J Antibiot (Tokyo) 1980;33:192–198. - PubMed
-
- Cassady JM, Chan KK, Floss HG, Leistner E. Chem Pharm Bull (Tokyo) 2004;52:1–26. - PubMed
-
- Spiteller P, Bai L, Shang G, Carroll BJ, Yu TW, Floss HG. J Am Chem Soc. 2003;125:14236–14237. - PubMed
-
- Xu J, Mahmud T, Floss HG. Arch Biochem Biophys. 2003;411:277–288. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

