In vivo genome editing restores haemostasis in a mouse model of haemophilia
- PMID: 21706032
- PMCID: PMC3152293
- DOI: 10.1038/nature10177
In vivo genome editing restores haemostasis in a mouse model of haemophilia
Abstract
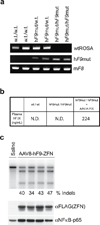
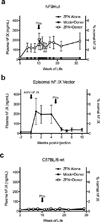
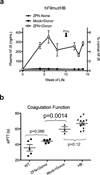
Editing of the human genome to correct disease-causing mutations is a promising approach for the treatment of genetic disorders. Genome editing improves on simple gene-replacement strategies by effecting in situ correction of a mutant gene, thus restoring normal gene function under the control of endogenous regulatory elements and reducing risks associated with random insertion into the genome. Gene-specific targeting has historically been limited to mouse embryonic stem cells. The development of zinc finger nucleases (ZFNs) has permitted efficient genome editing in transformed and primary cells that were previously thought to be intractable to such genetic manipulation. In vitro, ZFNs have been shown to promote efficient genome editing via homology-directed repair by inducing a site-specific double-strand break (DSB) at a target locus, but it is unclear whether ZFNs can induce DSBs and stimulate genome editing at a clinically meaningful level in vivo. Here we show that ZFNs are able to induce DSBs efficiently when delivered directly to mouse liver and that, when co-delivered with an appropriately designed gene-targeting vector, they can stimulate gene replacement through both homology-directed and homology-independent targeted gene insertion at the ZFN-specified locus. The level of gene targeting achieved was sufficient to correct the prolonged clotting times in a mouse model of haemophilia B, and remained persistent after induced liver regeneration. Thus, ZFN-driven gene correction can be achieved in vivo, raising the possibility of genome editing as a viable strategy for the treatment of genetic disease.
©2011 Macmillan Publishers Limited. All rights reserved
Figures


References
-
- Urnov FD, Rebar EJ, Holmes MC, Zhang HS, Gregory PD. Genome editing with engineered zinc finger nucleases. Nat Rev Genet. 2010;11(9):636–646. - PubMed
-
- Urnov FD, et al. Highly efficient endogenous human gene correction using designed zinc-finger nucleases. Nature. 2005;435(7042):646–651. - PubMed
-
- Porteus MH, Baltimore D. Chimeric nucleases stimulate gene targeting in human cells. Science. 2003;300(5620):763. - PubMed
-
- Cartier N, et al. Hematopoietic stem cell gene therapy with a lentiviral vector in X-linked adrenoleukodystrophy. Science. 2009;326(5954):818–823. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

