Two-step aminoacylation of tRNA without channeling in Archaea
- PMID: 21726564
- PMCID: PMC3156345
- DOI: 10.1016/j.jmb.2011.06.039
Two-step aminoacylation of tRNA without channeling in Archaea
Erratum in
- J Mol Biol. 2011 Oct 14;413(1):292-3
Abstract
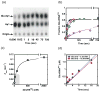
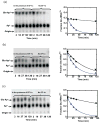
Catalysis of sequential reactions is often envisaged to occur by channeling of substrate between enzyme active sites without release into bulk solvent. However, while there are compelling physiological rationales for direct substrate transfer, proper experimental support for the hypothesis is often lacking, particularly for metabolic pathways involving RNA. Here, we apply transient kinetics approaches developed to study channeling in bienzyme complexes to an archaeal protein synthesis pathway featuring the misaminoacylated tRNA intermediate Glu-tRNA(Gln). Experimental and computational elucidation of a kinetic and thermodynamic framework for two-step cognate Gln-tRNA(Gln) synthesis demonstrates that the misacylating aminoacyl-tRNA synthetase (GluRS(ND)) and the tRNA-dependent amidotransferase (GatDE) function sequentially without channeling. Instead, rapid processing of the misacylated tRNA intermediate by GatDE and preferential elongation factor binding to the cognate Gln-tRNA(Gln) together permit accurate protein synthesis without formation of a binary protein-protein complex between GluRS(ND) and GatDE. These findings establish an alternate paradigm for protein quality control via two-step pathways for cognate aminoacyl-tRNA formation.
Copyright © 2011 Elsevier Ltd. All rights reserved.
Figures






Similar articles
-
The archaeal transamidosome for RNA-dependent glutamine biosynthesis.Nucleic Acids Res. 2010 Sep;38(17):5774-83. doi: 10.1093/nar/gkq336. Epub 2010 May 10. Nucleic Acids Res. 2010. PMID: 20457752 Free PMC article.
-
Structural basis for tRNA-dependent amidotransferase function.Structure. 2005 Oct;13(10):1421-33. doi: 10.1016/j.str.2005.06.016. Structure. 2005. PMID: 16216574
-
Two enzymes bound to one transfer RNA assume alternative conformations for consecutive reactions.Nature. 2010 Sep 30;467(7315):612-6. doi: 10.1038/nature09411. Nature. 2010. PMID: 20882017
-
Divergence of glutamate and glutamine aminoacylation pathways: providing the evolutionary rationale for mischarging.J Mol Evol. 1995 May;40(5):476-81. doi: 10.1007/BF00166615. J Mol Evol. 1995. PMID: 7783222 Review.
-
Evolution and variation in amide aminoacyl-tRNA synthesis.IUBMB Life. 2024 Aug;76(8):505-522. doi: 10.1002/iub.2811. Epub 2024 Feb 23. IUBMB Life. 2024. PMID: 38391119 Review.
Cited by
-
Yeast mitochondrial interactosome model: metabolon membrane proteins complex involved in the channeling of ADP/ATP.Int J Mol Sci. 2012;13(2):1858-1885. doi: 10.3390/ijms13021858. Epub 2012 Feb 10. Int J Mol Sci. 2012. PMID: 22408429 Free PMC article. Review.
-
A tRNA-independent mechanism for transamidosome assembly promotes aminoacyl-tRNA transamidation.J Biol Chem. 2013 Feb 8;288(6):3816-22. doi: 10.1074/jbc.M112.441394. Epub 2012 Dec 20. J Biol Chem. 2013. PMID: 23258533 Free PMC article.
-
Elements of the cellular metabolic structure.Front Mol Biosci. 2015 Apr 28;2:16. doi: 10.3389/fmolb.2015.00016. eCollection 2015. Front Mol Biosci. 2015. PMID: 25988183 Free PMC article.
-
Non-canonical roles of tRNAs and tRNA mimics in bacterial cell biology.Mol Microbiol. 2016 Aug;101(4):545-58. doi: 10.1111/mmi.13419. Epub 2016 Jun 28. Mol Microbiol. 2016. PMID: 27169680 Free PMC article. Review.
-
Lack of discrimination against non-proteinogenic amino acid norvaline by elongation factor Tu from Escherichia coli.Croat Chem Acta. 2013;86(1):73-82. doi: 10.5562/cca2173. Croat Chem Acta. 2013. PMID: 23750044 Free PMC article.
References
-
- Anderson KS. Fundamental mechanisms of substrate channeling. Methods Enzymol. 1999;308:111–126. - PubMed
-
- Huang X, Holden HM, Raushel FM. Channeling of substrates and intermediates in enzyme-catalyzed reactions. Annu Rev Biochem. 2001;70:149–180. - PubMed
-
- An S, Kumar R, Sheets ED, Benkovic SJ. Reversible compartmentalization of de novo purine biosynthetic complexes in living cells. Science. 2008;320:103–106. - PubMed
-
- Lenaz G, Genova ML. Structure and organization of mitochondrial respiratory complexes: a new understanding of an old subject. Antioxid Redox Signal. 2010;12:961–1008. - PubMed
-
- Meyer FM, Gerwig J, Hammer E, Herzberg C, Commichau FM, Volker U, Stulke J. Physical interactions between tricarboxylic acid cycle enzymes in Bacillus subtilis: evidence for a metabolon. Metab Eng. 2011;13:18–27. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

