Allostery in trypsin-like proteases suggests new therapeutic strategies
- PMID: 21726912
- PMCID: PMC3191250
- DOI: 10.1016/j.tibtech.2011.06.001
Allostery in trypsin-like proteases suggests new therapeutic strategies
Abstract
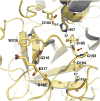
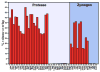
Trypsin-like proteases (TLPs) are a large family of enzymes responsible for digestion, blood coagulation, fibrinolysis, development, fertilization, apoptosis and immunity. A current paradigm posits that the irreversible transition from an inactive zymogen to the active protease form enables productive interaction with substrate and catalysis. Analysis of the entire structural database reveals two distinct conformations of the active site: one fully accessible to substrate (E) and the other occluded by the collapse of a specific segment (E*). The allosteric E*-E equilibrium provides a reversible mechanism for activity and regulation in addition to the irreversible zymogen to protease conversion and points to new therapeutic strategies aimed at inhibiting or activating the enzyme. In this review, we discuss relevant examples, with emphasis on the rational engineering of anticoagulant thrombin mutants.
Copyright © 2011 Elsevier Ltd. All rights reserved.
Figures

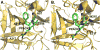

Similar articles
-
Crystallographic and kinetic evidence of allostery in a trypsin-like protease.Biochemistry. 2011 Jul 26;50(29):6301-7. doi: 10.1021/bi200878c. Epub 2011 Jun 30. Biochemistry. 2011. PMID: 21707111 Free PMC article.
-
Conformational selection in trypsin-like proteases.Curr Opin Struct Biol. 2012 Aug;22(4):421-31. doi: 10.1016/j.sbi.2012.05.006. Epub 2012 Jun 3. Curr Opin Struct Biol. 2012. PMID: 22664096 Free PMC article. Review.
-
Determinants of specificity in coagulation proteases.J Thromb Haemost. 2005 Nov;3(11):2401-8. doi: 10.1111/j.1538-7836.2005.01456.x. J Thromb Haemost. 2005. PMID: 16241939 Review.
-
Residue 225 determines the Na(+)-induced allosteric regulation of catalytic activity in serine proteases.Proc Natl Acad Sci U S A. 1996 Oct 1;93(20):10653-6. doi: 10.1073/pnas.93.20.10653. Proc Natl Acad Sci U S A. 1996. PMID: 8855234 Free PMC article.
-
Serine proteases.IUBMB Life. 2009 May;61(5):510-5. doi: 10.1002/iub.186. IUBMB Life. 2009. PMID: 19180666 Free PMC article. Review.
Cited by
-
Conformational Plasticity-Rigidity Axis of the Coagulation Factor VII Zymogen Elucidated by Atomistic Simulations of the N-Terminally Truncated Factor VIIa Protease Domain.Biomolecules. 2021 Apr 8;11(4):549. doi: 10.3390/biom11040549. Biomolecules. 2021. PMID: 33917935 Free PMC article.
-
The active site region plays a critical role in Na+ binding to thrombin.J Biol Chem. 2022 Jan;298(1):101458. doi: 10.1016/j.jbc.2021.101458. Epub 2021 Nov 30. J Biol Chem. 2022. PMID: 34861239 Free PMC article.
-
Pesticides as the drivers of neuropsychotic diseases, cancers, and teratogenicity among agro-workers as well as general public.Environ Sci Pollut Res Int. 2019 Jan;26(1):91-100. doi: 10.1007/s11356-018-3642-2. Epub 2018 Nov 8. Environ Sci Pollut Res Int. 2019. PMID: 30411285 Review.
-
Community Network Analysis of Allosteric Proteins.Methods Mol Biol. 2021;2253:137-151. doi: 10.1007/978-1-0716-1154-8_9. Methods Mol Biol. 2021. PMID: 33315222
-
Molecular mechanism of selective substrate engagement and inhibitor disengagement of cysteine synthase.J Biol Chem. 2021 Jan-Jun;296:100041. doi: 10.1074/jbc.RA120.014490. Epub 2020 Nov 24. J Biol Chem. 2021. PMID: 33162395 Free PMC article.
References
-
- Puente XS, Sanchez LM, Gutierrez-Fernandez A, Velasco G, Lopez-Otin C. A genomic view of the complexity of mammalian proteolytic systems. Biochem Soc Trans. 2005;33:331–334. - PubMed
-
- Ciechanover A, Iwai K. The ubiquitin system: from basic mechanisms to the patient bed. IUBMB Life. 2004;56:193–201. - PubMed
-
- Gomis-Ruth FX. Structural aspects of the metzincin clan of metalloendopeptidases. Mol Biotechnol. 2003;24:157–202. - PubMed
-
- Rea D, Fulop V. Structure-function properties of prolyl oligopeptidase family enzymes. Cell Biochem Biophys. 2006;44:349–365. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

