Amino-terminal extension present in the methionine aminopeptidase type 1c of Mycobacterium tuberculosis is indispensible for its activity
- PMID: 21729287
- PMCID: PMC3154147
- DOI: 10.1186/1471-2091-12-35
Amino-terminal extension present in the methionine aminopeptidase type 1c of Mycobacterium tuberculosis is indispensible for its activity
Abstract
Background: Methionine aminopeptidase (MetAP) is a ubiquitous enzyme in both prokaryotes and eukaryotes, which catalyzes co-translational removal of N-terminal methionine from elongating polypeptide chains during protein synthesis. It specifically removes the terminal methionine in all organisms, if the penultimate residue is non-bulky and uncharged. The MetAP action for exclusion of N-terminal methionine is mandatory in 50-70% of nascent proteins. Such an activity is required for proper sub cellular localization, additional processing and eventually for the degradation of proteins.
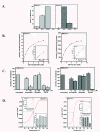
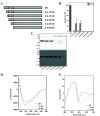
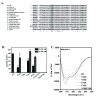
Results: We cloned genes encoding two such metalloproteases (MtMetAP1a and MtMetAP1c) present in Mycobacterium tuberculosis and expressed them as histidine-tagged proteins in Escherichia coli. Although they have different substrate preferences, for Met-Ala-Ser, we found, MtMetAP1c had significantly high enzyme turnover rate as opposed to MtMetAP1a. Circular dichroism spectroscopic studies as well as monitoring of enzyme activity indicated high temperature stability (up to 50 °C) of MtMetAP1a compared to that of the MtMetAP1c. Modelling of MtMetAP1a based on MtMetAP1c crystal structure revealed the distinct spatial arrangements of identical active site amino acid residues and their mutations affected the enzymatic activities of both the proteins. Strikingly, we observed that 40 amino acid long N-terminal extension of MtMetAP1c, compared to its other family members, contributes towards the activity and stability of this enzyme, which has never been reported for any methionine aminopeptidase. Furthermore, mutational analysis revealed that Val-18 and Pro-19 of MtMetAP1c are crucial for its enzymatic activity. Consistent with this observation, molecular dynamic simulation studies of wild-type and these variants strongly suggest their involvement in maintaining active site conformation of MtMetAP1c.
Conclusion: Our findings unequivocally emphasized that N-terminal extension of MtMetAP1c contributes towards the functionality of the enzyme presumably by regulating active site residues through "action-at-a-distance" mechanism and we for the first time are reporting this unique function of the enzyme.
Figures


Similar articles
-
Anti-Tuberculosis Potential of OJT008 against Active and Multi-Drug-Resistant Mycobacterium Tuberculosis: In Silico and In Vitro Inhibition of Methionine Aminopeptidase.Int J Mol Sci. 2023 Dec 5;24(24):17142. doi: 10.3390/ijms242417142. Int J Mol Sci. 2023. PMID: 38138972 Free PMC article.
-
Catalysis and inhibition of Mycobacterium tuberculosis methionine aminopeptidase.J Med Chem. 2010 Feb 11;53(3):1329-37. doi: 10.1021/jm901624n. J Med Chem. 2010. PMID: 20038112 Free PMC article.
-
Expression and characterization of two functional methionine aminopeptidases from Mycobacterium tuberculosis H37Rv.Curr Microbiol. 2009 Nov;59(5):520-5. doi: 10.1007/s00284-009-9470-3. Epub 2009 Aug 18. Curr Microbiol. 2009. PMID: 19688379
-
Structure and function of the methionine aminopeptidases.Biochim Biophys Acta. 2000 Mar 7;1477(1-2):157-67. doi: 10.1016/s0167-4838(99)00271-x. Biochim Biophys Acta. 2000. PMID: 10708856 Review.
-
Methionine aminopeptidases and angiogenesis.Essays Biochem. 2002;38:65-78. doi: 10.1042/bse0380065. Essays Biochem. 2002. PMID: 12463162 Review.
Cited by
-
MapB Protein is the Essential Methionine Aminopeptidase in Mycobacterium tuberculosis.Cells. 2019 Apr 28;8(5):393. doi: 10.3390/cells8050393. Cells. 2019. PMID: 31035386 Free PMC article.
-
Anti-Tuberculosis Potential of OJT008 against Active and Multi-Drug-Resistant Mycobacterium Tuberculosis: In Silico and In Vitro Inhibition of Methionine Aminopeptidase.Int J Mol Sci. 2023 Dec 5;24(24):17142. doi: 10.3390/ijms242417142. Int J Mol Sci. 2023. PMID: 38138972 Free PMC article.
-
Biochemical characterization of recombinant methionine aminopeptidases (MAPs) from Mycobacterium tuberculosis H37Rv.Mol Cell Biochem. 2012 Jun;365(1-2):191-202. doi: 10.1007/s11010-012-1260-8. Epub 2012 Apr 1. Mol Cell Biochem. 2012. PMID: 22466806
-
Purification of native HBHA from Mycobacterium avium subsp. paratuberculosis.BMC Res Notes. 2013 Feb 7;6:55. doi: 10.1186/1756-0500-6-55. BMC Res Notes. 2013. PMID: 23390963 Free PMC article.
-
Fragment-Sized Thiazoles in Fragment-Based Drug Discovery Campaigns: Friend or Foe?ACS Med Chem Lett. 2022 Nov 3;13(12):1905-1910. doi: 10.1021/acsmedchemlett.2c00429. eCollection 2022 Dec 8. ACS Med Chem Lett. 2022. PMID: 36518695 Free PMC article.
References
Publication types
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources
Miscellaneous

