Microbial Communities as Experimental Units
- PMID: 21731083
- PMCID: PMC3128510
- DOI: 10.1525/bio.2011.61.5.9
Microbial Communities as Experimental Units
Abstract
Artificial ecosystem selection is an experimental technique that treats microbial communities as though they were discrete units by applying selection on community-level properties. Highly diverse microbial communities associated with humans and other organisms can have significant impacts on the health of the host. It is difficult to find correlations between microbial community composition and community-associated diseases, in part because it may be impossible to define a universal and robust species concept for microbes. Microbial communities are composed of potentially thousands of unique populations that evolved in intimate contact, so it is appropriate in many situations to view the community as the unit of analysis. This perspective is supported by recent discoveries using metagenomics and pangenomics. Artificial ecosystem selection experiments can be costly, but they bring the logical rigor of biological model systems to the emerging field of microbial community analysis.
Figures



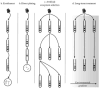
References
-
- Bashey F, Lively CM. Group selection on population size affects life-history patterns in the entomopathogenic nematode Steinernema carpocapsae. Evolution. 2009;63:1301–1311. - PubMed
-
- Bateson G. Mind and Nature: A Necessary Unity. E P Dutton; 1979.
-
- Brenner K, You L, Arnold F. Engineering microbial consortia: A new frontier in synthetic biology. Trends in Biotechnology. 2008;26:483–489. - PubMed
-
- Dinsdale EA, et al. Functional metagenomic profiling of nine biomes. Nature. 2008;452:629–632. - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources

