Robust network topologies for generating switch-like cellular responses
- PMID: 21731481
- PMCID: PMC3121696
- DOI: 10.1371/journal.pcbi.1002085
Robust network topologies for generating switch-like cellular responses
Abstract
Signaling networks that convert graded stimuli into binary, all-or-none cellular responses are critical in processes ranging from cell-cycle control to lineage commitment. To exhaustively enumerate topologies that exhibit this switch-like behavior, we simulated all possible two- and three-component networks on random parameter sets, and assessed the resulting response profiles for both steepness (ultrasensitivity) and extent of memory (bistability). Simulations were used to study purely enzymatic networks, purely transcriptional networks, and hybrid enzymatic/transcriptional networks, and the topologies in each class were rank ordered by parametric robustness (i.e., the percentage of applied parameter sets exhibiting ultrasensitivity or bistability). Results reveal that the distribution of network robustness is highly skewed, with the most robust topologies clustering into a small number of motifs. Hybrid networks are the most robust in generating ultrasensitivity (up to 28%) and bistability (up to 18%); strikingly, a purely transcriptional framework is the most fragile in generating either ultrasensitive (up to 3%) or bistable (up to 1%) responses. The disparity in robustness among the network classes is due in part to zero-order ultrasensitivity, an enzyme-specific phenomenon, which repeatedly emerges as a particularly robust mechanism for generating nonlinearity and can act as a building block for switch-like responses. We also highlight experimentally studied examples of topologies enabling switching behavior, in both native and synthetic systems, that rank highly in our simulations. This unbiased approach for identifying topologies capable of a given response may be useful in discovering new natural motifs and in designing robust synthetic gene networks.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures



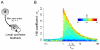
 in that parameter set (where b is the basal synthesis rate and v is the maximal feedback synthesis rate). If
in that parameter set (where b is the basal synthesis rate and v is the maximal feedback synthesis rate). If  is sufficiently high, then the Hill coefficient reaches a maximum when the effective feedback synthesis rate constant
is sufficiently high, then the Hill coefficient reaches a maximum when the effective feedback synthesis rate constant  (where KF is the threshold concentration) is approximately equal to the degradation rate constant kdeg.
(where KF is the threshold concentration) is approximately equal to the degradation rate constant kdeg.


References
-
- Alon U. Network motifs: theory and experimental approaches. Nat Rev Genet. 2007;8:450–461. doi: 10.1038/nrg2102. - DOI - PubMed
-
- Breitkreutz A, Choi H, Sharom JR, Boucher L, Neduva V, et al. A Global Protein Kinase and Phosphatase Interaction Network in Yeast. Science. 2010;328:1043–1046. doi: 10.1126/science.1176495. - DOI - PMC - PubMed
-
- Ferrell JE. Feedback regulation of opposing enzymes generates robust, all-or-none bistable responses. Curr Biol. 2008;18:R244–R245. doi: 10.1016/j.cub.2008.02.035. - DOI - PMC - PubMed
-
- Malleshaiah MK, Shahrezaei V, Swain PS, Michnick SW. The scaffold protein Ste5 directly controls a switch-like mating decision in yeast. Nature. 2010;465:101–105. doi: 10.1038/nature08946. - DOI - PubMed
-
- Xiong W, Ferrell JE. A positive-feedback-based bistable ‘memory module’ that governs a cell fate decision. Nature. 2003;426:460–465. doi: 10.1038/nature02089. - DOI - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

