Crystallizing Membrane Proteins in Lipidic Mesophases. A Host Lipid Screen
- PMID: 21743796
- PMCID: PMC3131202
- DOI: 10.1021/cg101378s
Crystallizing Membrane Proteins in Lipidic Mesophases. A Host Lipid Screen
Abstract
The default lipid for the bulk of the crystallogenesis studies performed to date using the cubic mesophase method is monoolein. There is no good reason however, why this 18-carbon, cis-monounsaturated monoacylglycerol should be the preferred lipid for all target membrane proteins. The latter come from an array of biomembrane types with varying properties that include hydrophobic thickness, intrinsic curvature, lateral pressure profile, lipid and protein makeup, and compositional asymmetry. Thus, it seems reasonable that screening for crystallizability based on the identity of the lipid creating the hosting mesophase would be worthwhile. For this, monoacylglycerols with differing acyl chain characteristics, such as length and olefinic bond position, must be available. A lipid synthesis and purification program is in place in the author's laboratory to serve this need. In the current study with the outer membrane sugar transporter, OprB, we demonstrate the utility of host lipid screening as a means for generating diffraction-quality crystals. Host lipid screening is likely to prove a generally useful strategy for mesophase-based crystallization of membrane proteins.
Figures



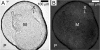



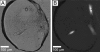

Similar articles
-
Crystallizing Membrane Proteins in the Lipidic Mesophase. Experience with Human Prostaglandin E2 Synthase 1 and an Evolving Strategy.Cryst Growth Des. 2014 Apr 2;14(4):2034-2047. doi: 10.1021/cg500157x. Epub 2014 Mar 7. Cryst Growth Des. 2014. PMID: 24803849 Free PMC article.
-
Host Lipid and Temperature as Important Screening Variables for Crystallizing Integral Membrane Proteins in Lipidic Mesophases. Trials with Diacylglycerol Kinase.Cryst Growth Des. 2013 Jul 3;13(7):2846-2857. doi: 10.1021/cg400254v. Cryst Growth Des. 2013. PMID: 23956688 Free PMC article.
-
Membrane protein crystallization in lipidic mesophases with tailored bilayers.Structure. 2004 Dec;12(12):2113-24. doi: 10.1016/j.str.2004.09.020. Structure. 2004. PMID: 15576026
-
A comprehensive review of the lipid cubic phase or in meso method for crystallizing membrane and soluble proteins and complexes.Acta Crystallogr F Struct Biol Commun. 2015 Jan 1;71(Pt 1):3-18. doi: 10.1107/S2053230X14026843. Epub 2015 Jan 1. Acta Crystallogr F Struct Biol Commun. 2015. PMID: 25615961 Free PMC article. Review.
-
Crystallizing membrane proteins for structure determination: use of lipidic mesophases.Annu Rev Biophys. 2009;38:29-51. doi: 10.1146/annurev.biophys.050708.133655. Annu Rev Biophys. 2009. PMID: 19086821 Review.
Cited by
-
Crystallizing membrane proteins for structure-function studies using lipidic mesophases.Biochem Soc Trans. 2011 Jun;39(3):725-32. doi: 10.1042/BST0390725. Biochem Soc Trans. 2011. PMID: 21599641 Free PMC article.
-
Preparation of 1-Monoacylglycerols via the Suzuki-Miyaura Reaction: 2,3-Dihydroxypropyl (Z)-tetradec-7-enoate.Organic Synth. 2012;89:183-201. doi: 10.1002/0471264229.os089.19. Organic Synth. 2012. PMID: 25505351 Free PMC article. No abstract available.
-
Lipidic cubic phase technologies for membrane protein structural studies.Curr Opin Struct Biol. 2011 Aug;21(4):559-66. doi: 10.1016/j.sbi.2011.06.007. Epub 2011 Jul 19. Curr Opin Struct Biol. 2011. PMID: 21775127 Free PMC article. Review.
-
In situ flow cell platform for examining calcium oxalate and calcium phosphate crystallization on films of basement membrane extract in the presence of urinary 'inhibitors'.CrystEngComm. 2020 Feb 28;22(8):1448-1458. doi: 10.1039/c9ce01587f. Epub 2020 Feb 5. CrystEngComm. 2020. PMID: 32256199 Free PMC article.
-
Crystallizing Membrane Proteins in the Lipidic Mesophase. Experience with Human Prostaglandin E2 Synthase 1 and an Evolving Strategy.Cryst Growth Des. 2014 Apr 2;14(4):2034-2047. doi: 10.1021/cg500157x. Epub 2014 Mar 7. Cryst Growth Des. 2014. PMID: 24803849 Free PMC article.
References
Grants and funding
LinkOut - more resources
Full Text Sources
