Comprehensive exon array data processing method for quantitative analysis of alternative spliced variants
- PMID: 21745820
- PMCID: PMC3185423
- DOI: 10.1093/nar/gkr513
Comprehensive exon array data processing method for quantitative analysis of alternative spliced variants
Abstract
Alternative splicing of pre-mRNA generates protein diversity. Dysfunction of splicing machinery and expression of specific transcripts has been linked to cancer progression and drug response. Exon microarray technology enables genome-wide quantification of expression levels of the majority of exons and facilitates the discovery of alternative splicing events. Analysis of exon array data is more challenging than the analysis of gene expression data and there is a need for reliable quantification of exons and alternatively spliced variants. We introduce a novel, computationally efficient methodology, Multiple Exon Array Preprocessing (MEAP), for exon array data pre-processing, analysis and visualization. We compared MEAP with existing pre-processing methods, and validation of six exons and two alternatively spliced variants with qPCR corroborated MEAP expression estimates. Analysis of exon array data from head and neck squamous cell carcinoma (HNSCC) cell lines revealed several transcripts associated with 11q13 amplification, which is related with decreased survival and metastasis in HNSCC patients. Our results demonstrate that MEAP produces reliable expression values at exon, alternatively spliced variant and gene levels, which allows generating novel experimentally testable predictions.
Figures

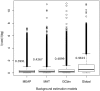


 ), fold change (
), fold change ( ), expression intensities of each exon in a gene between two sample groups (Group1: 11q13+; Group2: 11q13−). Gaps may occur when there are no probes designed against a specific exon region. Sections `Expression' and `Transcript' give the exon expression profile and the expression of each splice variant of ORAOV1 in 11q13+/11q13−. G1/G2 in the exon expression profile corresponds to the fold change between 11q13+ and 11q13− samples, in which green represents downregulation and red represents upregulation. Expression values are on
), expression intensities of each exon in a gene between two sample groups (Group1: 11q13+; Group2: 11q13−). Gaps may occur when there are no probes designed against a specific exon region. Sections `Expression' and `Transcript' give the exon expression profile and the expression of each splice variant of ORAOV1 in 11q13+/11q13−. G1/G2 in the exon expression profile corresponds to the fold change between 11q13+ and 11q13− samples, in which green represents downregulation and red represents upregulation. Expression values are on  -scale.
-scale.
Similar articles
-
A statistical framework for genome-wide discovery of biomarker splice variations with GeneChip Human Exon 1.0 ST Arrays.Genome Inform. 2006;17(1):88-99. Genome Inform. 2006. PMID: 17503359
-
Expression microarray analysis reveals alternative splicing of LAMA3 and DST genes in head and neck squamous cell carcinoma.PLoS One. 2014 Mar 27;9(3):e91263. doi: 10.1371/journal.pone.0091263. eCollection 2014. PLoS One. 2014. PMID: 24675808 Free PMC article.
-
Alternative splicing and differential gene expression in colon cancer detected by a whole genome exon array.BMC Genomics. 2006 Dec 27;7:325. doi: 10.1186/1471-2164-7-325. BMC Genomics. 2006. PMID: 17192196 Free PMC article.
-
Functional regulation of alternatively spliced Na+/Ca2+ exchanger (NCX1) isoforms.Ann N Y Acad Sci. 2002 Nov;976:187-96. doi: 10.1111/j.1749-6632.2002.tb04740.x. Ann N Y Acad Sci. 2002. PMID: 12502560 Review.
-
Sequence and Evolutionary Features for the Alternatively Spliced Exons of Eukaryotic Genes.Int J Mol Sci. 2019 Aug 6;20(15):3834. doi: 10.3390/ijms20153834. Int J Mol Sci. 2019. PMID: 31390737 Free PMC article. Review.
Cited by
-
Aberrant expression of CPSF1 promotes head and neck squamous cell carcinoma via regulating alternative splicing.PLoS One. 2020 May 21;15(5):e0233380. doi: 10.1371/journal.pone.0233380. eCollection 2020. PLoS One. 2020. PMID: 32437477 Free PMC article.
-
puma 3.0: improved uncertainty propagation methods for gene and transcript expression analysis.BMC Bioinformatics. 2013 Feb 5;14:39. doi: 10.1186/1471-2105-14-39. BMC Bioinformatics. 2013. PMID: 23379655 Free PMC article.
-
Deregulation of COMMD1 is associated with poor prognosis in diffuse large B-cell lymphoma.PLoS One. 2014 Mar 13;9(3):e91031. doi: 10.1371/journal.pone.0091031. eCollection 2014. PLoS One. 2014. PMID: 24625556 Free PMC article. Clinical Trial.
-
Alternative splicing discriminates molecular subtypes and has prognostic impact in diffuse large B-cell lymphoma.Blood Cancer J. 2017 Aug 25;7(8):e596. doi: 10.1038/bcj.2017.71. Blood Cancer J. 2017. PMID: 28841210 Free PMC article. Clinical Trial.
-
Netrin-4 promotes glioblastoma cell proliferation through integrin β4 signaling.Neoplasia. 2012 Mar;14(3):219-27. doi: 10.1593/neo.111396. Neoplasia. 2012. PMID: 22496621 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

