Targeted Sos1 deletion reveals its critical role in early T-cell development
- PMID: 21746917
- PMCID: PMC3145744
- DOI: 10.1073/pnas.1104295108
Targeted Sos1 deletion reveals its critical role in early T-cell development
Abstract
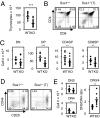
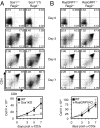
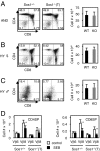
Activation of the small G protein Ras is required for thymocyte differentiation. In thymocytes, Ras is activated by the Ras guanine exchange factors (RasGEFs) Sos1, Sos2, and RasGRP1. We report the development of a floxed allele of sos1 to assess the role of Sos1 during thymocyte development. Sos1 was required for pre-T-cell receptor (pre-TCR)- but not TCR-stimulated developmental signals. Sos1 deletion led to a partial block at the DN-to-DP transition. Sos1-deficient thymocytes showed reduced pre-TCR-stimulated proliferation, differentiation, and ERK phosphorylation. In contrast, TCR-stimulated positive selection, and negative selection under strong stimulatory conditions, remained intact in Sos1-deficient mice. Comparison of RasGEF expression at different developmental stages showed that relative to Sos2 and RasGRP1, Sos1 is most abundant in DN thymocytes, but least abundant in DP thymocytes. These data reveal that Sos1 is uniquely positioned to affect signal transduction early in thymocyte development.
Conflict of interest statement
The authors declare no conflict of interest.
Figures


References
-
- Buday L, Downward J. Many faces of Ras activation. Biochim Biophys Acta. 2008;1786:178–187. - PubMed
-
- Hanahan D, Weinberg RA. The hallmarks of cancer. Cell. 2000;100:57–70. - PubMed
-
- Mor A, Philips MR. Compartmentalized Ras/MAPK signaling. Annu Rev Immunol. 2006;24:771–800. - PubMed
-
- Daniels MA, et al. Thymic selection threshold defined by compartmentalization of Ras/MAPK signalling. Nature. 2006;444:724–729. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

