Improving the performance of β-turn prediction using predicted shape strings and a two-layer support vector machine model
- PMID: 21749732
- PMCID: PMC3155507
- DOI: 10.1186/1471-2105-12-283
Improving the performance of β-turn prediction using predicted shape strings and a two-layer support vector machine model
Abstract
Background: The β-turn is a secondary protein structure type that plays an important role in protein configuration and function. Development of accurate prediction methods to identify β-turns in protein sequences is valuable. Several methods for β-turn prediction have been developed; however, the prediction quality is still a challenge and there is substantial room for improvement. Innovations of the proposed method focus on discovering effective features, and constructing a new architectural model.
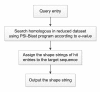
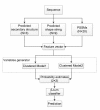
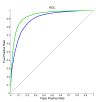
Results: We utilized predicted secondary structures, predicted shape strings and the position-specific scoring matrix (PSSM) as input features, and proposed a novel two-layer model to enhance the prediction. We achieved the highest values according to four evaluation measures, i.e. Q(total) = 87.2%, MCC = 0.66, Q(observed) = 75.9%, and Q(predicted) = 73.8% on the BT426 dataset. The results show that our proposed two-layer model discriminates better between β-turns and non-β-turns than the single model due to obtaining higher Q(predicted). Moreover, the predicted shape strings based on the structural alignment approach greatly improve the performance, and the same improvements were observed on BT547 and BT823 datasets as well.
Conclusion: In this article, we present a comprehensive method for the prediction of β-turns. Experiments show that the proposed method constitutes a great improvement over the competing prediction methods.
Figures
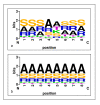
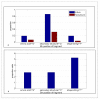
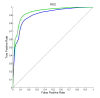
References
-
- Richardson JS. The anatomy and taxonomy of protein structure. Adv Protein Chem. 1981;34:167–339. - PubMed
-
- Rose GD, Gierasch LM, Smith JA. Turns in peptides and proteins. Adv Protein Chem. 1985;37:1–109. - PubMed
-
- Guruprasad K, Rajkumar S. Beta-and gamma-turns in proteins revisited: a new set of amino acid turn-type dependent positional preferences and potentials. J Biosci. 2000;25(2):143–156. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

