Parallel evolution of transcriptome architecture during genome reorganization
- PMID: 21750103
- PMCID: PMC3205574
- DOI: 10.1101/gr.122218.111
Parallel evolution of transcriptome architecture during genome reorganization
Abstract
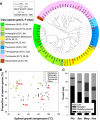
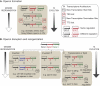
Assembly of genes into operons is generally viewed as an important process during the continual adaptation of microbes to changing environmental challenges. However, the genome reorganization events that drive this process are also the roots of instability for existing operons. We have determined that there exists a statistically significant trend that correlates the proportion of genes encoded in operons in archaea to their phylogenetic lineage. We have further characterized how microbes deal with operon instability by mapping and comparing transcriptome architectures of four phylogenetically diverse extremophiles that span the range of operon stabilities observed across archaeal lineages: a photoheterotrophic halophile (Halobacterium salinarum NRC-1), a hydrogenotrophic methanogen (Methanococcus maripaludis S2), an acidophilic and aerobic thermophile (Sulfolobus solfataricus P2), and an anaerobic hyperthermophile (Pyrococcus furiosus DSM 3638). We demonstrate how the evolution of transcriptional elements (promoters and terminators) generates new operons, restores the coordinated regulation of translocated, inverted, and newly acquired genes, and introduces completely novel regulation for even some of the most conserved operonic genes such as those encoding subunits of the ribosome. The inverse correlation (r=-0.92) between the proportion of operons with such internally located transcriptional elements and the fraction of conserved operons in each of the four archaea reveals an unprecedented view into varying stages of operon evolution. Importantly, our integrated analysis has revealed that organisms adapted to higher growth temperatures have lower tolerance for genome reorganization events that disrupt operon structures.
Figures




References
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
