Population genetics of Glossina palpalis palpalis from central African sleeping sickness foci
- PMID: 21767402
- PMCID: PMC3162924
- DOI: 10.1186/1756-3305-4-140
Population genetics of Glossina palpalis palpalis from central African sleeping sickness foci
Abstract
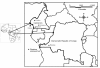
Background: Glossina palpalis palpalis (Diptera: Glossinidae) is widespread in west Africa, and is the main vector of sleeping sickness in Cameroon as well as in the Bas Congo Province of the Democratic Republic of Congo. However, little is known on the structure of its populations. We investigated G. p. palpalis population genetic structure in five sleeping sickness foci (four in Cameroon, one in Democratic Republic of Congo) using eight microsatellite DNA markers.
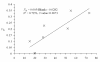
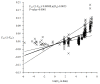
Results: A strong isolation by distance explains most of the population structure observed in our sampling sites of Cameroon and DRC. The populations here are composed of panmictic subpopulations occupying fairly wide zones with a very strong isolation by distance. Effective population sizes are probably between 20 and 300 individuals and if we assume densities between 120 and 2000 individuals per km2, dispersal distance between reproducing adults and their parents extends between 60 and 300 meters.
Conclusions: This first investigation of population genetic structure of G. p. palpalis in Central Africa has evidenced random mating subpopulations over fairly large areas and is thus at variance with that found in West African populations of G. p. palpalis. This study brings new information on the isolation by distance at a macrogeographic scale which in turn brings useful information on how to organise regional tsetse control. Future investigations should be directed at temporal sampling to have more accurate measures of demographic parameters in order to help vector control decision.
Figures

Similar articles
-
Single-strand conformation polymorphism (SSCP) of mitochondrial genes helps to estimate genetic differentiation, demographic parameters and phylogeny of Glossina palpalis palpalis populations from West and Central Africa.Infect Genet Evol. 2020 Aug;82:104303. doi: 10.1016/j.meegid.2020.104303. Epub 2020 Apr 3. Infect Genet Evol. 2020. PMID: 32247869
-
Ecotype evolution in Glossina palpalis subspecies, major vectors of sleeping sickness.PLoS Negl Trop Dis. 2015 Mar 16;9(3):e0003497. doi: 10.1371/journal.pntd.0003497. eCollection 2015 Mar. PLoS Negl Trop Dis. 2015. PMID: 25775377 Free PMC article.
-
Population genetics of Glossina palpalis palpalis in sleeping sickness foci of Côte d'Ivoire before and after vector control.Infect Genet Evol. 2019 Nov;75:103963. doi: 10.1016/j.meegid.2019.103963. Epub 2019 Jul 10. Infect Genet Evol. 2019. PMID: 31301424 Free PMC article.
-
How can tsetse population genetics contribute to African trypanosomiasis control?Trends Parasitol. 2010 May;26(5):255-63. doi: 10.1016/j.pt.2010.02.006. Epub 2010 Mar 2. Trends Parasitol. 2010. PMID: 20202905 Review.
-
[Population growth and global warming: impacts on tsetse and trypanosomoses in West Africa].Parasite. 2009 Mar;16(1):3-10. doi: 10.1051/parasite/2009161003. Parasite. 2009. PMID: 19353946 Review. French.
Cited by
-
Spatial meta-analysis of the occurrence and distribution of tsetse-transmitted animal trypanosomiasis in Cameroon over the last 30 years.Epidemiol Infect. 2022 Apr 27;150:1-38. doi: 10.1017/S0950268822000772. Online ahead of print. Epidemiol Infect. 2022. PMID: 35473820 Free PMC article.
-
Intestinal Bacterial Communities of Trypanosome-Infected and Uninfected Glossina palpalis palpalis from Three Human African Trypanomiasis Foci in Cameroon.Front Microbiol. 2017 Aug 3;8:1464. doi: 10.3389/fmicb.2017.01464. eCollection 2017. Front Microbiol. 2017. PMID: 28824591 Free PMC article.
-
Detection of Wolbachia and different trypanosome species in Glossina palpalis palpalis populations from three sleeping sickness foci of southern Cameroon.Parasit Vectors. 2018 Dec 12;11(1):630. doi: 10.1186/s13071-018-3229-2. Parasit Vectors. 2018. PMID: 30541614 Free PMC article.
-
Challenges towards the elimination of Human African Trypanosomiasis in the sleeping sickness focus of Campo in southern Cameroon.Parasit Vectors. 2014 Aug 16;7:374. doi: 10.1186/1756-3305-7-374. Parasit Vectors. 2014. PMID: 25129168 Free PMC article. Review.
-
Association between IL1 gene polymorphism and human African trypanosomiasis in populations of sleeping sickness foci of southern Cameroon.PLoS Negl Trop Dis. 2019 Mar 25;13(3):e0007283. doi: 10.1371/journal.pntd.0007283. eCollection 2019 Mar. PLoS Negl Trop Dis. 2019. PMID: 30908482 Free PMC article.
References
-
- Simarro P, Cecchi G, Paone M, Franco JR, Diarra A, Ruiz Postigo JR, Fèvre EM, Courtin F, Mattioli RC, Jannin JG. The Atlas of human African trypanosomiasis: a contribution to global mapping of neglected tropical diseases. Int J Health Geogr. 2010;9:57. doi: 10.1186/1476-072X-9-57. - DOI - PMC - PubMed
-
- Dyer NA, Lawton SP, Ravel S, Choi KS, Lehane MJ, Robinson AS, Okedi LA, Hall M, Solano P, Donnelly MJ. Molecular phylogenetics of tsetse flies (Diptera: Glossinidae) based on mitochondrial (CO1, 16S, ND2) and nuclear ribosomal DNA sequences, with an emphasis on the palpalis group. Mol Phylog Evol. 2008;49:227–239. doi: 10.1016/j.ympev.2008.07.011. - DOI - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources

