SPR-5 is a histone H3K4 demethylase with a role in meiotic double-strand break repair
- PMID: 21768382
- PMCID: PMC3150895
- DOI: 10.1073/pnas.1102298108
SPR-5 is a histone H3K4 demethylase with a role in meiotic double-strand break repair
Abstract
Regulation of histone methylation levels has long been implicated in multiple cellular processes, many of which involve transcription. Here, however, we report a unique role for the Caenorhabditis elegans histone demethylase SPR-5 in meiotic DNA double-strand break repair (DSBR). SPR-5 shows enzymatic activity toward H3K4me2 both in vitro and in the nematode germline, and spr-5 mutants show several phenotypes indicating a perturbation of DSBR, including increased p53-dependent germ cell apoptosis, increased levels of the DSBR marker RAD-51, and sensitivity toward DSB-inducing treatments. spr-5 mutants show no transcriptional misregulation of known DSBR involved genes. Instead, SPR-5 shows a rapid subcellular relocalization upon DSB-inducing treatment, which suggests that SPR-5 may function directly in DSBR.
Conflict of interest statement
The authors declare no conflict of interest.
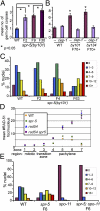
Figures




Similar articles
-
C. elegans germ cells switch between distinct modes of double-strand break repair during meiotic prophase progression.PLoS Genet. 2007 Nov;3(11):e191. doi: 10.1371/journal.pgen.0030191. PLoS Genet. 2007. PMID: 17983271 Free PMC article.
-
ZTF-8 interacts with the 9-1-1 complex and is required for DNA damage response and double-strand break repair in the C. elegans germline.PLoS Genet. 2014 Oct 16;10(10):e1004723. doi: 10.1371/journal.pgen.1004723. eCollection 2014 Oct. PLoS Genet. 2014. PMID: 25329393 Free PMC article.
-
Fanconi Anemia FANCM/FNCM-1 and FANCD2/FCD-2 Are Required for Maintaining Histone Methylation Levels and Interact with the Histone Demethylase LSD1/SPR-5 in Caenorhabditis elegans.Genetics. 2018 Jun;209(2):409-423. doi: 10.1534/genetics.118.300823. Epub 2018 Mar 27. Genetics. 2018. PMID: 29588287 Free PMC article.
-
Reversal of histone lysine trimethylation by the JMJD2 family of histone demethylases.Cell. 2006 May 5;125(3):467-81. doi: 10.1016/j.cell.2006.03.028. Epub 2006 Apr 6. Cell. 2006. PMID: 16603238
-
LET-418/Mi2 and SPR-5/LSD1 cooperatively prevent somatic reprogramming of C. elegans germline stem cells.Stem Cell Reports. 2014 Mar 27;2(4):547-59. doi: 10.1016/j.stemcr.2014.02.007. eCollection 2014 Apr 8. Stem Cell Reports. 2014. PMID: 24749077 Free PMC article.
Cited by
-
LSD1-LIKE1-Mediated H3K4me2 Demethylation Is Required for Homologous Recombination Repair.Plant Physiol. 2019 Oct;181(2):499-509. doi: 10.1104/pp.19.00530. Epub 2019 Jul 31. Plant Physiol. 2019. PMID: 31366719 Free PMC article.
-
H3K4 Methylation in Aging and Metabolism.Epigenomes. 2021 Jun 18;5(2):14. doi: 10.3390/epigenomes5020014. Epigenomes. 2021. PMID: 34968301 Free PMC article. Review.
-
Triptolide induces cell-cycle arrest and apoptosis of human multiple myeloma cells in vitro via altering expression of histone demethylase LSD1 and JMJD2B.Acta Pharmacol Sin. 2012 Jan;33(1):109-19. doi: 10.1038/aps.2011.145. Epub 2011 Nov 28. Acta Pharmacol Sin. 2012. PMID: 22120968 Free PMC article.
-
H3K4 demethylase activities repress proliferative and postmitotic aging.Aging Cell. 2014 Apr;13(2):245-53. doi: 10.1111/acel.12166. Epub 2013 Nov 19. Aging Cell. 2014. PMID: 24134677 Free PMC article.
-
Kinetic analysis of strand invasion during C. elegans meiosis reveals similar rates of sister- and homolog-directed repair.bioRxiv [Preprint]. 2025 Jan 10:2025.01.10.632442. doi: 10.1101/2025.01.10.632442. bioRxiv. 2025. PMID: 39829846 Free PMC article. Preprint.
References
-
- Martin C, Zhang Y. The diverse functions of histone lysine methylation. Nat Rev Mol Cell Biol. 2005;6:838–849. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

