Identity by descent estimation with dense genome-wide genotype data
- PMID: 21769932
- PMCID: PMC3587128
- DOI: 10.1002/gepi.20606
Identity by descent estimation with dense genome-wide genotype data
Abstract
We present a novel method, IBDLD, for estimating the probability of identity by descent (IBD) for a pair of related individuals at a locus, given dense genotype data and a pedigree of arbitrary size and complexity. IBDLD overcomes the challenges of exact multipoint estimation of IBD in pedigrees of potentially large size and eliminates the difficulty of accommodating the background linkage disequilibrium (LD) that is present in high-density genotype data. We show that IBDLD is much more accurate at estimating the true IBD sharing than methods that remove LD by pruning SNPs and is highly robust to pedigree errors or other forms of misspecified relationships. The method is fast and can be used to estimate the probability for each possible IBD sharing state at every SNP from a high-density genotyping array for hundreds of thousands of pairs of individuals. We use it to estimate point-wise and genomewide IBD sharing between 185,745 pairs of subjects all of whom are related through a single, large and complex 13-generation pedigree and genotyped with the Affymetrix 500 k chip. We find that we are able to identify the true pedigree relationship for individuals who were misidentified in the collected data and estimate empirical kinship coefficients that can be used in follow-up QTL mapping studies. IBDLD is implemented as an open source software package and is freely available.
© 2011 Wiley-Liss, Inc.
Figures
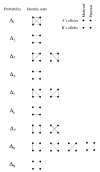
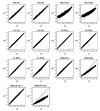
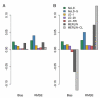
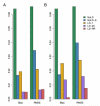

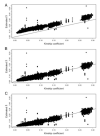

References
-
- Abecasis GR, Cherny SS, Cookson WO, Cardon LR. Merlin–rapid analysis of dense genetic maps using sparse gene flow trees. Nat Genet. 2002;30:97–101. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

