The subcellular distribution of small molecules: a meta-analysis
- PMID: 21774504
- PMCID: PMC3185174
- DOI: 10.1021/mp200093z
The subcellular distribution of small molecules: a meta-analysis
Abstract
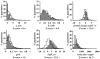
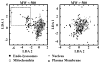
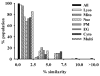
To explore the extent to which current knowledge about the organelle-targeting features of small molecules may be applicable toward controlling the accumulation and distribution of exogenous chemical agents inside cells, molecules with known subcellular localization properties (as reported in the scientific literature) were compiled into a single data set. This data set was compared to a reference data set of approved drug molecules derived from the DrugBank database, and to a reference data set of random organic molecules derived from the PubChem database. Cheminformatic analysis revealed that molecules with reported subcellular localizations were comparably diverse. However, the calculated physicochemical properties of molecules reported to accumulate in different organelles were markedly overlapping. In relation to the reference sets of DrugBank and PubChem molecules, molecules with reported subcellular localizations were biased toward larger, more complex chemical structures possessing multiple ionizable functional groups and higher lipophilicity. Stratifying molecules based on molecular weight revealed that many physicochemical properties' trends associated with specific organelles were reversed in smaller vs larger molecules. Most likely, these reversed trends are due to the different transport mechanisms determining the subcellular localization of molecules of different sizes. Molecular weight can be dramatically altered by tagging molecules with fluorophores or by incorporating organelle targeting motifs. Generally, in order to better exploit structure-localization relationships, subcellular targeting strategies would benefit from analysis of the biodistribution effects resulting from variations in the size of the molecules.
Figures



Similar articles
-
The subcellular distribution of small molecules: from pharmacokinetics to synthetic biology.Mol Pharm. 2011 Oct 3;8(5):1619-28. doi: 10.1021/mp200092v. Epub 2011 Aug 15. Mol Pharm. 2011. PMID: 21805990 Free PMC article. Review.
-
Drug-likeness and increased hydrophobicity of commercially available compound libraries for drug screening.Curr Top Med Chem. 2012;12(14):1500-13. doi: 10.2174/156802612802652466. Curr Top Med Chem. 2012. PMID: 22827520 Review.
-
Guiding Drugs to Target-Harboring Organelles: Stretching Drug-Delivery to a Higher Level of Resolution.Angew Chem Int Ed Engl. 2019 Oct 28;58(44):15584-15594. doi: 10.1002/anie.201906284. Epub 2019 Aug 12. Angew Chem Int Ed Engl. 2019. PMID: 31237741 Review.
-
[Strategy of molecular design of drugs: the unification of macro-properties and micro-structures of a molecule].Yao Xue Xue Bao. 2008 Mar;43(3):227-33. Yao Xue Xue Bao. 2008. PMID: 18630256 Chinese.
-
Evolution of the physicochemical properties of marketed drugs: can history foretell the future?Drug Discov Today. 2011 Nov;16(21-22):976-84. doi: 10.1016/j.drudis.2011.07.003. Epub 2011 Jul 18. Drug Discov Today. 2011. PMID: 21782967 Review.
Cited by
-
Development of background-free tame fluorescent probes for intracellular live cell imaging.Nat Commun. 2016 Jun 20;7:11964. doi: 10.1038/ncomms11964. Nat Commun. 2016. PMID: 27321135 Free PMC article.
-
Structurally Redesigned Bioorthogonal Reagents for Mitochondria-Specific Prodrug Activation.JACS Au. 2021 Jan 25;1(1):23-30. doi: 10.1021/jacsau.0c00053. Epub 2020 Dec 15. JACS Au. 2021. PMID: 33554213 Free PMC article.
-
Labeling Preferences of Diazirines with Protein Biomolecules.J Am Chem Soc. 2021 May 5;143(17):6691-6700. doi: 10.1021/jacs.1c02509. Epub 2021 Apr 20. J Am Chem Soc. 2021. PMID: 33876925 Free PMC article.
-
Visualizing chemical structure-subcellular localization relationships using fluorescent small molecules as probes of cellular transport.J Cheminform. 2013 Oct 5;5(1):44. doi: 10.1186/1758-2946-5-44. J Cheminform. 2013. PMID: 24093553 Free PMC article.
-
Nanoscale chemical imaging using tip-enhanced Raman spectroscopy.Nat Protoc. 2019 Apr;14(4):1169-1193. doi: 10.1038/s41596-019-0132-z. Epub 2019 Mar 25. Nat Protoc. 2019. PMID: 30911174
References
-
- Rosania GR. Supertargeted chemistry: identifying relationships between molecular structures and their sub-cellular distribution. Curr Top Med Chem. 2003;3:659–85. - PubMed
-
- Lipinski CA, Lombardo F, Dominy BW, Feeney PJ. Experimental and computational approaches to estimate solubility and permeability in drug discovery and development settings. Adv Drug Deliv Rev. 2001;46:3–26. - PubMed
-
- Feng MR. Assessment of blood-brain barrier penetration: in silico, in vitro and in vivo. Curr Drug Metab. 2002;3:647–57. - PubMed
-
- Rodgers T, Leahy D, Rowland M. Physiologically based pharmacokinetic modeling 1: predicting the tissue distribution of moderate-to-strong bases. J Pharm Sci. 2005;94:1259–76. - PubMed
-
- Rodgers T, Rowland M. Physiologically based pharmacokinetic modelling 2: predicting the tissue distribution of acids, very weak bases, neutrals and zwitterions. J Pharm Sci. 2006;95:1238–57. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

