Genome-wide association study identifies novel restless legs syndrome susceptibility loci on 2p14 and 16q12.1
- PMID: 21779176
- PMCID: PMC3136436
- DOI: 10.1371/journal.pgen.1002171
Genome-wide association study identifies novel restless legs syndrome susceptibility loci on 2p14 and 16q12.1
Erratum in
- PLoS Genet. 2011 Aug;7(8). doi: 10.1371/annotation/393ad2d3-df4f-4770-87bc-00bfabf79362
Abstract
Restless legs syndrome (RLS) is a sensorimotor disorder with an age-dependent prevalence of up to 10% in the general population above 65 years of age. Affected individuals suffer from uncomfortable sensations and an urge to move in the lower limbs that occurs mainly in resting situations during the evening or at night. Moving the legs or walking leads to an improvement of symptoms. Concomitantly, patients report sleep disturbances with consequences such as reduced daytime functioning. We conducted a genome-wide association study (GWA) for RLS in 922 cases and 1,526 controls (using 301,406 SNPs) followed by a replication of 76 candidate SNPs in 3,935 cases and 5,754 controls, all of European ancestry. Herein, we identified six RLS susceptibility loci of genome-wide significance, two of them novel: an intergenic region on chromosome 2p14 (rs6747972, P = 9.03 × 10(-11), OR = 1.23) and a locus on 16q12.1 (rs3104767, P = 9.4 × 10(-19), OR = 1.35) in a linkage disequilibrium block of 140 kb containing the 5'-end of TOX3 and the adjacent non-coding RNA BC034767.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures


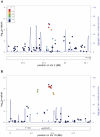
References
-
- Allen RP, Picchietti D, Hening WA, Trenkwalder C, Walters AS, et al. Restless legs syndrome: diagnostic criteria, special considerations, and epidemiology. A report from the restless legs syndrome diagnosis and epidemiology workshop at the National Institutes of Health. Sleep Med. 2003;4:101–119. - PubMed
-
- Winkelmann J, Schormair B, Lichtner P, Ripke S, Xiong L, et al. Genome-wide association study of restless legs syndrome identifies common variants in three genomic regions. Nat Genet. 2007;39:1000–1006. - PubMed
-
- Stefansson H, Rye DB, Hicks A, Petursson H, Ingason A, et al. A genetic risk factor for periodic limb movements in sleep. N Engl J Med. 2007;357:639–647. - PubMed
-
- Schormair B, Kemlink D, Roeske D, Eckstein G, Xiong L, et al. PTPRD (protein tyrosine phosphatase receptor type delta) is associated with restless legs syndrome. Nat Genet. 2008;40:946–948. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

