PwHAP5, a CCAAT-binding transcription factor, interacts with PwFKBP12 and plays a role in pollen tube growth orientation in Picea wilsonii
- PMID: 21784992
- PMCID: PMC3192995
- DOI: 10.1093/jxb/err120
PwHAP5, a CCAAT-binding transcription factor, interacts with PwFKBP12 and plays a role in pollen tube growth orientation in Picea wilsonii
Abstract
The HAP complex occurs in many eukaryotic organisms and is involved in multiple physiological processes. Here it was found that in Picea wilsonii, HAP5 (PwHAP5), a putative CCAAT-binding transcription factor gene, is involved in pollen tube development and control of tube orientation. Quantitative real-time reverse transcription-PCR showed that PwHAP5 transcripts were expressed strongly in germinating pollen and could be induced by Ca(2+). Overexpression of PwHAP5 in pollen altered pollen tube orientation, whereas the tube with PwHAP5RNAi showed normal growth without diminishing pollen tube growth. Furthermore, PwFKBP12, which encodes an FK506-binding protein (FKBP) was screened and a bimolecular fluorescence complementation assay performed to confirm the interaction of PwHAP5 and PwFKBP12 in vivo. Transient expression of PwFKBP12 in pollen showed normal pollen tube growth, whereas the tube with PwFKBP12RNAi bent. The phenotype of overexpression of HAP5 on pollen tube was restored by FKBP12. Altogether, our study supported the role of HAP5 in pollen tube development and orientation regulation and identified FKBP12 as a novel partner to interact with HAP5 involved in the process.
Figures


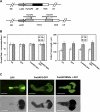



References
-
- Ben-Naim O, Eshed R, Parnis A, Teper-Bamnolker P, Shalit A, Coupland G, Samach A, Lifschitz E. The CCAAT binding factor can mediate interactions between CONSTANS-like proteins and DNA. The Plant Journal. 2006;46:462–476. - PubMed
-
- Bucher P. Weight matrix descriptions of four eukaryotic RNA polymerase II promoter elements derived from 502 unrelated promoter sequences. Journal of Molecular Biology. 1990;212:563–578. - PubMed
-
- Chang S, Purgear J, Cairney J. A simple and efficient method for isolating RNA from pine trees. Plant Molecular Biology Reporter. 1993;11:117–121.
-
- Chen NZ, Zhang XQ, Wei PC, Chen QJ, Ren F, Chen J, Wang XC. AtHAP3b plays a crucial role in the regulation of flowering time in Arabidopsis during osmotic stress. Journal of Biochemistry and Molecular Biology. 2007;40:1083–1089. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources
Miscellaneous

