Evidence for network evolution in an Arabidopsis interactome map
- PMID: 21798944
- PMCID: PMC3170756
- DOI: 10.1126/science.1203877
Evidence for network evolution in an Arabidopsis interactome map
Abstract
Plants have unique features that evolved in response to their environments and ecosystems. A full account of the complex cellular networks that underlie plant-specific functions is still missing. We describe a proteome-wide binary protein-protein interaction map for the interactome network of the plant Arabidopsis thaliana containing about 6200 highly reliable interactions between about 2700 proteins. A global organization of plant biological processes emerges from community analyses of the resulting network, together with large numbers of novel hypothetical functional links between proteins and pathways. We observe a dynamic rewiring of interactions following gene duplication events, providing evidence for a model of evolution acting upon interactome networks. This and future plant interactome maps should facilitate systems approaches to better understand plant biology and improve crops.
Figures
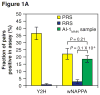









Comment in
-
Cell biology. A cellular roadmap for the plant kingdom.Science. 2011 Jul 29;333(6042):532-3. doi: 10.1126/science.1209753. Science. 2011. PMID: 21798921 No abstract available.
-
Technology: Getting Moore from DNA sequencing.Nat Rev Genet. 2011 Aug 2;12(9):586. doi: 10.1038/nrg3059. Nat Rev Genet. 2011. PMID: 21808262 No abstract available.
-
Systems biology: Plant networks.Nat Rev Genet. 2011 Aug 9;12(9):586. doi: 10.1038/nrg3058. Nat Rev Genet. 2011. PMID: 21826065 No abstract available.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

