Comprehensive, atomic-level characterization of structurally characterized protein-protein interactions: the PICCOLO database
- PMID: 21801404
- PMCID: PMC3161047
- DOI: 10.1186/1471-2105-12-313
Comprehensive, atomic-level characterization of structurally characterized protein-protein interactions: the PICCOLO database
Abstract
Background: Structural studies are increasingly providing huge amounts of information on multi-protein assemblies. Although a complete understanding of cellular processes will be dependent on an explicit characterization of the intermolecular interactions that underlie these assemblies and mediate molecular recognition, these are not well described by standard representations.
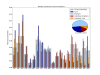
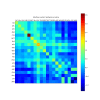
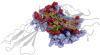
Results: Here we present PICCOLO, a comprehensive relational database capturing the details of structurally characterized protein-protein interactions. Interactions are described at the level of interacting pairs of atoms, residues and polypeptide chains, with the physico-chemical nature of the interactions being characterized. Distance and angle terms are used to distinguish 12 different interaction types, including van der Waals contacts, hydrogen bonds and hydrophobic contacts. The explicit aim of PICCOLO is to underpin large-scale analyses of the properties of protein-protein interfaces. This is exemplified by an analysis of residue propensity and interface contact preferences derived from a much larger data set than previously reported. However, PICCOLO also supports detailed inspection of particular systems of interest.
Conclusions: The current PICCOLO database comprises more than 260 million interacting atom pairs from 38,202 protein complexes. A web interface for the database is available at http://www-cryst.bioc.cam.ac.uk/piccolo.
Figures
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

