The origin of patterning systems in bilateria-insights from the Hox and ParaHox genes in Acoelomorpha
- PMID: 21802044
- PMCID: PMC5054442
- DOI: 10.1016/S1672-0229(11)60010-7
The origin of patterning systems in bilateria-insights from the Hox and ParaHox genes in Acoelomorpha
Abstract
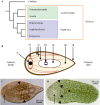
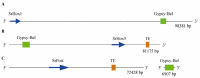
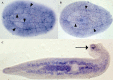
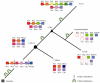
Hox and ParaHox genes constitute two families of developmental regulators that pattern the Anterior-Posterior body axis in all bilaterians. The members of these two groups of genes are usually arranged in genomic clusters and work in a coordinated fashion, both in space and in time. While the mechanistic aspects of their action are relatively well known, it is still unclear how these systems evolved. For instance, we still need a proper model of how the Hox and ParaHox clusters were assembled over time. This problem is due to the shortage of information on gene complements for many taxa (mainly basal metazoans) and the lack of a consensus phylogenetic model of animal relationships to which we can relate our new findings. Recently, several studies have shown that the Acoelomorpha most probably represent the first offshoot of the Bilateria. This finding has prompted us, and others, to study the Hox and ParaHox complements in these animals, as well as their activity during development. In this review, we analyze how the current knowledge of Hox and ParaHox genes in the Acoelomorpha is shaping our view of bilaterian evolution.
Copyright © 2011 Beijing Genomics Institute. Published by Elsevier Ltd. All rights reserved.
Figures
References
-
- McGinnis W., Krumlauf R. Homeobox genes and axial patterning. Cell. 1992;68:283–302. - PubMed
-
- Bürglin T.R. Homeodomain proteins. In: Meyers R.A., editor. Vol.3. VCH Verlagsgesellschaft mbH; Weinheim, Germany: 1996. pp. 55–76. (Encyclopedia of Molecular Biology and Molecular Medicine).
-
- Quiquand M. More constraint on ParaHox than Hox gene families in early metazoan evolution. Dev. Biol. 2009;328:173–187. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

