Arabidopsis mutants deleted in the light-harvesting protein Lhcb4 have a disrupted photosystem II macrostructure and are defective in photoprotection
- PMID: 21803939
- PMCID: PMC3226214
- DOI: 10.1105/tpc.111.087320
Arabidopsis mutants deleted in the light-harvesting protein Lhcb4 have a disrupted photosystem II macrostructure and are defective in photoprotection
Abstract
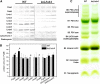
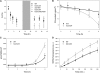
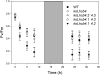
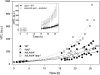
The role of the light-harvesting complex Lhcb4 (CP29) in photosynthesis was investigated in Arabidopsis thaliana by characterizing knockout lines for each of the three Lhcb4 isoforms (Lhcb4.1/4.2/4.3). Plants lacking all isoforms (koLhcb4) showed a compensatory increase of Lhcb1 and a slightly reduced photosystem II/I ratio with respect to the wild type. The absence of Lhcb4 did not result in alteration in electron transport rates. However, the kinetic of state transition was faster in the mutant, and nonphotochemical quenching activity was lower in koLhcb4 plants with respect to either wild type or mutants retaining a single Lhcb4 isoform. KoLhcb4 plants were more sensitive to photoinhibition, while this effect was not observed in knockout lines for any other photosystem II antenna subunit. Ultrastructural analysis of thylakoid grana membranes showed a lower density of photosystem II complexes in koLhcb4. Moreover, analysis of isolated supercomplexes showed a different overall shape of the C₂S₂ particles due to a different binding mode of the S-trimer to the core complex. An empty space was observed within the photosystem II supercomplex at the Lhcb4 position, implying that the missing Lhcb4 was not replaced by other Lhc subunits. This suggests that Lhcb4 is unique among photosystem II antenna proteins and determinant for photosystem II macro-organization and photoprotection.
Figures




Similar articles
-
The roles of Arabidopsis proteins of Lhcb4, Lhcb5 and Lhcb6 in oxidative stress under natural light conditions.Plant Physiol Biochem. 2018 Sep;130:267-276. doi: 10.1016/j.plaphy.2018.07.014. Epub 2018 Jul 17. Plant Physiol Biochem. 2018. PMID: 30032070
-
Disturbed excitation energy transfer in Arabidopsis thaliana mutants lacking minor antenna complexes of photosystem II.Biochim Biophys Acta. 2014 Dec;1837(12):1981-1988. doi: 10.1016/j.bbabio.2014.09.011. Biochim Biophys Acta. 2014. PMID: 25291424
-
Towards spruce-type photosystem II: consequences of the loss of light-harvesting proteins LHCB3 and LHCB6 in Arabidopsis.Plant Physiol. 2021 Dec 4;187(4):2691-2715. doi: 10.1093/plphys/kiab396. Plant Physiol. 2021. PMID: 34618099 Free PMC article.
-
LHC-like Proteins: The Guardians of Photosynthesis.Int J Mol Sci. 2023 Jan 28;24(3):2503. doi: 10.3390/ijms24032503. Int J Mol Sci. 2023. PMID: 36768826 Free PMC article. Review.
-
Phylogenetic viewpoints on regulation of light harvesting and electron transport in eukaryotic photosynthetic organisms.Planta. 2013 Feb;237(2):399-412. doi: 10.1007/s00425-012-1744-5. Epub 2012 Sep 13. Planta. 2013. PMID: 22971817 Review.
Cited by
-
The Arabidopsis Proteins AtNHR2A and AtNHR2B Are Multi-Functional Proteins Integrating Plant Immunity With Other Biological Processes.Front Plant Sci. 2020 Mar 4;11:232. doi: 10.3389/fpls.2020.00232. eCollection 2020. Front Plant Sci. 2020. PMID: 32194606 Free PMC article.
-
Quenching in Arabidopsis thaliana mutants lacking monomeric antenna proteins of photosystem II.J Biol Chem. 2011 Oct 21;286(42):36830-40. doi: 10.1074/jbc.M111.273227. Epub 2011 Aug 15. J Biol Chem. 2011. PMID: 21844190 Free PMC article.
-
Salicylic Acid Protects Photosystem II by Alleviating Photoinhibition in Arabidopsis thaliana under High Light.Int J Mol Sci. 2020 Feb 12;21(4):1229. doi: 10.3390/ijms21041229. Int J Mol Sci. 2020. PMID: 32059402 Free PMC article.
-
Light Harvesting in Fluctuating Environments: Evolution and Function of Antenna Proteins across Photosynthetic Lineage.Plants (Basel). 2021 Jun 10;10(6):1184. doi: 10.3390/plants10061184. Plants (Basel). 2021. PMID: 34200788 Free PMC article.
-
Phylogenetic, Structural and Functional Evolution of the LHC Gene Family in Plant Species.Int J Mol Sci. 2022 Dec 28;24(1):488. doi: 10.3390/ijms24010488. Int J Mol Sci. 2022. PMID: 36613939 Free PMC article.
References
-
- Ahn T.K., Avenson T.J., Ballottari M., Cheng Y.C., Niyogi K.K., Bassi R., Fleming G.R. (2008). Architecture of a charge-transfer state regulating light harvesting in a plant antenna protein. Science 320: 794–797 - PubMed
-
- Alboresi A., Dall’osto L., Aprile A., Carillo P., Roncaglia E., Cattivelli L., Bassi R. (2011). Reactive oxygen species and transcript analysis upon excess light treatment in wild-type Arabidopsis thaliana vs a photosensitive mutant lacking zeaxanthin and lutein. BMC Plant Biol. 11: 62. - PMC - PubMed
-
- Allen J.F. (1992). Protein phosphorylation in regulation of photosynthesis. Biochim. Biophys. Acta 1098: 275–335 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

