Synergistic use of plant-prokaryote comparative genomics for functional annotations
- PMID: 21810204
- PMCID: PMC3223725
- DOI: 10.1186/1471-2164-12-S1-S2
Synergistic use of plant-prokaryote comparative genomics for functional annotations
Abstract
Background: Identifying functions for all gene products in all sequenced organisms is a central challenge of the post-genomic era. However, at least 30-50% of the proteins encoded by any given genome are of unknown or vaguely known function, and a large number are wrongly annotated. Many of these 'unknown' proteins are common to prokaryotes and plants. We set out to predict and experimentally test the functions of such proteins. Our approach to functional prediction integrates comparative genomics based mainly on microbial genomes with functional genomic data from model microorganisms and post-genomic data from plants. This approach bridges the gap between automated homology-based annotations and the classical gene discovery efforts of experimentalists, and is more powerful than purely computational approaches to identifying gene-function associations.
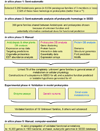
Results: Among Arabidopsis genes, we focused on those (2,325 in total) that (i) are unique or belong to families with no more than three members, (ii) occur in prokaryotes, and (iii) have unknown or poorly known functions. Computer-assisted selection of promising targets for deeper analysis was based on homology-independent characteristics associated in the SEED database with the prokaryotic members of each family. In-depth comparative genomic analysis was performed for 360 top candidate families. From this pool, 78 families were connected to general areas of metabolism and, of these families, specific functional predictions were made for 41. Twenty-one predicted functions have been experimentally tested or are currently under investigation by our group in at least one prokaryotic organism (nine of them have been validated, four invalidated, and eight are in progress). Ten additional predictions have been independently validated by other groups. Discovering the function of very widespread but hitherto enigmatic proteins such as the YrdC or YgfZ families illustrates the power of our approach.
Conclusions: Our approach correctly predicted functions for 19 uncharacterized protein families from plants and prokaryotes; none of these functions had previously been correctly predicted by computational methods. The resulting annotations could be propagated with confidence to over six thousand homologous proteins encoded in over 900 bacterial, archaeal, and eukaryotic genomes currently available in public databases.
Figures

Similar articles
-
An Experimental Approach to Genome Annotation: This report is based on a colloquium sponsored by the American Academy of Microbiology held July 19-20, 2004, in Washington, DC.Washington (DC): American Society for Microbiology; 2004. Washington (DC): American Society for Microbiology; 2004. PMID: 33001599 Free Books & Documents. Review.
-
Genome-wide analysis of CCCH zinc finger family in Arabidopsis and rice.BMC Genomics. 2008 Jan 27;9:44. doi: 10.1186/1471-2164-9-44. BMC Genomics. 2008. PMID: 18221561 Free PMC article.
-
GreenPhylDB v5: a comparative pangenomic database for plant genomes.Nucleic Acids Res. 2021 Jan 8;49(D1):D1464-D1471. doi: 10.1093/nar/gkaa1068. Nucleic Acids Res. 2021. PMID: 33237299 Free PMC article.
-
Genome-wide analysis of the rice and Arabidopsis non-specific lipid transfer protein (nsLtp) gene families and identification of wheat nsLtp genes by EST data mining.BMC Genomics. 2008 Feb 21;9:86. doi: 10.1186/1471-2164-9-86. BMC Genomics. 2008. PMID: 18291034 Free PMC article.
-
Comparative genomics approaches to understanding and manipulating plant metabolism.Curr Opin Biotechnol. 2013 Apr;24(2):278-84. doi: 10.1016/j.copbio.2012.07.005. Epub 2012 Aug 14. Curr Opin Biotechnol. 2013. PMID: 22898705 Review.
Cited by
-
The MORPH algorithm: ranking candidate genes for membership in Arabidopsis and tomato pathways.Plant Cell. 2012 Nov;24(11):4389-406. doi: 10.1105/tpc.112.104513. Epub 2012 Nov 30. Plant Cell. 2012. PMID: 23204403 Free PMC article.
-
Metabolite damage and its repair or pre-emption.Nat Chem Biol. 2013 Feb;9(2):72-80. doi: 10.1038/nchembio.1141. Nat Chem Biol. 2013. PMID: 23334546 Review.
-
Systematic discovery of novel eukaryotic transcriptional regulators using sequence homology independent prediction.BMC Genomics. 2017 Jun 26;18(1):480. doi: 10.1186/s12864-017-3853-9. BMC Genomics. 2017. PMID: 28651538 Free PMC article.
-
Chronic industrial perturbation and seasonal change induces shift in the bacterial community from gammaproteobacteria to betaproteobacteria having catabolic potential for aromatic compounds at Amlakhadi canal.World J Microbiol Biotechnol. 2023 Dec 26;40(2):52. doi: 10.1007/s11274-023-03848-1. World J Microbiol Biotechnol. 2023. PMID: 38146029
-
A Comparative Metagenomic Analysis of Specified Microorganisms in Groundwater for Non-Sterilized Pharmaceutical Products.Curr Microbiol. 2024 Jul 17;81(9):273. doi: 10.1007/s00284-024-03791-w. Curr Microbiol. 2024. PMID: 39017960 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

