Regulatory cross-talk links Vibrio cholerae chromosome II replication and segregation
- PMID: 21811418
- PMCID: PMC3141006
- DOI: 10.1371/journal.pgen.1002189
Regulatory cross-talk links Vibrio cholerae chromosome II replication and segregation
Abstract
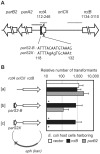
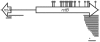
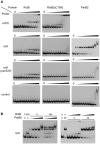
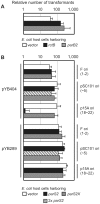
There is little knowledge of factors and mechanisms for coordinating bacterial chromosome replication and segregation. Previous studies have revealed that genes (and their products) that surround the origin of replication (oriCII) of Vibrio cholerae chromosome II (chrII) are critical for controlling the replication and segregation of this chromosome. rctB, which flanks one side of oriCII, encodes a protein that initiates chrII replication; rctA, which flanks the other side of oriCII, inhibits rctB activity. The chrII parAB2 operon, which is essential for chrII partitioning, is located immediately downstream of rctA. Here, we explored how rctA exerts negative control over chrII replication. Our observations suggest that RctB has at least two DNA binding domains--one for binding to oriCII and initiating replication and the other for binding to rctA and thereby inhibiting RctB's ability to initiate replication. Notably, the inhibitory effect of rctA could be alleviated by binding of ParB2 to a centromere-like parS site within rctA. Furthermore, by binding to rctA, ParB2 and RctB inversely regulate expression of the parAB2 genes. Together, our findings suggest that fluctuations in binding of the partitioning protein ParB2 and the chrII initiator RctB to rctA underlie a regulatory network controlling both oriCII firing and the production of the essential chrII partitioning proteins. Thus, by binding both RctB and ParB2, rctA serves as a nexus for regulatory cross-talk coordinating chrII replication and segregation.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures

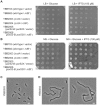
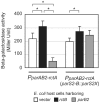
Similar articles
-
Molecular Dissection of the Essential Features of the Origin of Replication of the Second Vibrio cholerae Chromosome.mBio. 2015 Jul 28;6(4):e00973. doi: 10.1128/mBio.00973-15. mBio. 2015. PMID: 26220967 Free PMC article.
-
Chromosome I controls chromosome II replication in Vibrio cholerae.PLoS Genet. 2014 Feb 27;10(2):e1004184. doi: 10.1371/journal.pgen.1004184. eCollection 2014 Feb. PLoS Genet. 2014. PMID: 24586205 Free PMC article.
-
Transcriptional inactivation of a regulatory site for replication of Vibrio cholerae chromosome II.Proc Natl Acad Sci U S A. 2006 Aug 8;103(32):12051-6. doi: 10.1073/pnas.0605120103. Epub 2006 Jul 27. Proc Natl Acad Sci U S A. 2006. PMID: 16873545 Free PMC article.
-
Management of multipartite genomes: the Vibrio cholerae model.Curr Opin Microbiol. 2014 Dec;22:120-6. doi: 10.1016/j.mib.2014.10.003. Curr Opin Microbiol. 2014. PMID: 25460805 Review.
-
Cell cycle-coordinated maintenance of the Vibrio bipartite genome.EcoSal Plus. 2023 Dec 12;11(1):eesp00082022. doi: 10.1128/ecosalplus.esp-0008-2022. Epub 2023 Nov 22. EcoSal Plus. 2023. PMID: 38277776 Free PMC article. Review.
Cited by
-
Chromosome dynamics in multichromosome bacteria.Biochim Biophys Acta. 2012 Jul;1819(7):826-9. doi: 10.1016/j.bbagrm.2012.01.012. Epub 2012 Jan 28. Biochim Biophys Acta. 2012. PMID: 22306663 Free PMC article. Review.
-
Characterization of chromosomal and megaplasmid partitioning loci in Thermus thermophilus HB27.BMC Genomics. 2015 Apr 18;16(1):317. doi: 10.1186/s12864-015-1523-3. BMC Genomics. 2015. PMID: 25909452 Free PMC article.
-
Initiator protein dimerization plays a key role in replication control of Vibrio cholerae chromosome 2.Nucleic Acids Res. 2014;42(16):10538-49. doi: 10.1093/nar/gku771. Epub 2014 Aug 26. Nucleic Acids Res. 2014. PMID: 25159619 Free PMC article.
-
Chromosome segregation proteins of Vibrio cholerae as transcription regulators.mBio. 2014 May 6;5(3):e01061-14. doi: 10.1128/mBio.01061-14. mBio. 2014. PMID: 24803519 Free PMC article.
-
Molecular Dissection of the Essential Features of the Origin of Replication of the Second Vibrio cholerae Chromosome.mBio. 2015 Jul 28;6(4):e00973. doi: 10.1128/mBio.00973-15. mBio. 2015. PMID: 26220967 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

