Handpicking epigenetic marks with PHD fingers
- PMID: 21813457
- PMCID: PMC3241642
- DOI: 10.1093/nar/gkr613
Handpicking epigenetic marks with PHD fingers
Abstract
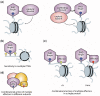
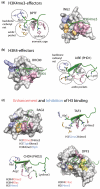
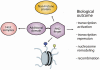
Plant homeodomain (PHD) fingers have emerged as one of the largest families of epigenetic effectors capable of recognizing or 'reading' post-translational histone modifications and unmodified histone tails. These interactions are highly specific and can be modulated by the neighboring epigenetic marks and adjacent effectors. A few PHD fingers have recently been found to also associate with non-histone proteins. In this review, we detail the molecular mechanisms and biological outcomes of the histone and non-histone targeting by PHD fingers. We discuss the significance of crosstalk between the histone modifications and consequences of combinatorial readout for selective recruitment of the PHD finger-containing components of chromatin remodeling and transcriptional complexes.
Figures
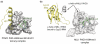
References
-
- Luger K, Mader AW, Richmond RK, Sargent DF, Richmond TJ. Crystal structure of the nucleosome core particle at 2.8 A resolution. Nature. 1997;389:251–260. - PubMed
-
- Strahl BD, Allis CD. The language of covalent histone modifications. Nature. 2000;403:41–45. - PubMed
-
- Jenuwein T, Allis CD. Translating the histone code. Science. 2001;293:1074–1080. - PubMed
-
- Turner BM. Cellular memory and the histone code. Cell. 2002;111:285–291. - PubMed
-
- Kouzarides T. Chromatin modifications and their function. Cell. 2007;128:693–705. - PubMed

