Crystal structures of enterovirus 71 3C protease complexed with rupintrivir reveal the roles of catalytically important residues
- PMID: 21813612
- PMCID: PMC3196404
- DOI: 10.1128/JVI.05107-11
Crystal structures of enterovirus 71 3C protease complexed with rupintrivir reveal the roles of catalytically important residues
Abstract
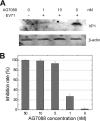
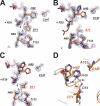
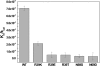
EV71 is the primary pathogenic cause of hand-foot-mouth disease (HFMD), but an effective antiviral drug currently is unavailable. Rupintrivir, an inhibitor against human rhinovirus (HRV), has potent antiviral activities against EV71. We determined the high-resolution crystal structures of the EV71 3C(pro)/rupintrivir complex, showing that although rupintrivir interacts with EV71 3C(pro) similarly to HRV 3C(pro), the C terminus of the inhibitor cannot accommodate the leaving-group pockets of EV71 3C(pro). Our structures reveal that EV71 3C(pro) possesses a surface-recessive S2' pocket that is not present in HRV 3C(pro) that contributes to the additional substrate binding affinity. Combined with mutagenic studies, we demonstrated that catalytic Glu71 is irreplaceable for maintaining the overall architecture of the active site and, most importantly, the productive conformation of catalytic His40. We discovered the role of a previously uncharacterized residue, Arg39 of EV71 3C(pro), that can neutralize the negative charge of Glu71, which may subsequently assist deprotonation of His40 during proteolysis.
Figures



Similar articles
-
Enterovirus 71 and coxsackievirus A16 3C proteases: binding to rupintrivir and their substrates and anti-hand, foot, and mouth disease virus drug design.J Virol. 2011 Oct;85(19):10319-31. doi: 10.1128/JVI.00787-11. Epub 2011 Jul 27. J Virol. 2011. PMID: 21795339 Free PMC article.
-
Structures of Enterovirus 71 3C proteinase (strain E2004104-TW-CDC) and its complex with rupintrivir.Acta Crystallogr D Biol Crystallogr. 2013 May;69(Pt 5):866-71. doi: 10.1107/S0907444913002862. Epub 2013 Apr 19. Acta Crystallogr D Biol Crystallogr. 2013. PMID: 23633597
-
Structural basis for antiviral inhibition of the main protease, 3C, from human enterovirus 93.J Virol. 2011 Oct;85(20):10764-73. doi: 10.1128/JVI.05062-11. Epub 2011 Aug 10. J Virol. 2011. PMID: 21835784 Free PMC article.
-
The Function and Mechanism of Enterovirus 71 (EV71) 3C Protease.Curr Microbiol. 2020 Sep;77(9):1968-1975. doi: 10.1007/s00284-020-02082-4. Epub 2020 Jun 15. Curr Microbiol. 2020. PMID: 32556480 Review.
-
Structural analysis of foot-and-mouth disease virus 3C protease: a viable target for antiviral drugs?Biochem Soc Trans. 2007 Jun;35(Pt 3):594-8. doi: 10.1042/BST0350594. Biochem Soc Trans. 2007. PMID: 17511659 Review.
Cited by
-
Enterovirus A71 antivirals: Past, present, and future.Acta Pharm Sin B. 2022 Apr;12(4):1542-1566. doi: 10.1016/j.apsb.2021.08.017. Epub 2021 Aug 20. Acta Pharm Sin B. 2022. PMID: 35847514 Free PMC article.
-
Enterovirus 71 3C inhibits cytokine expression through cleavage of the TAK1/TAB1/TAB2/TAB3 complex.J Virol. 2014 Sep 1;88(17):9830-41. doi: 10.1128/JVI.01425-14. Epub 2014 Jun 18. J Virol. 2014. PMID: 24942571 Free PMC article.
-
Molecular Evolution of the Protease Region in Norovirus Genogroup II.Front Microbiol. 2020 Jan 14;10:2991. doi: 10.3389/fmicb.2019.02991. eCollection 2019. Front Microbiol. 2020. PMID: 31993031 Free PMC article.
-
Structure of Main Protease from Human Coronavirus NL63: Insights for Wide Spectrum Anti-Coronavirus Drug Design.Sci Rep. 2016 Mar 7;6:22677. doi: 10.1038/srep22677. Sci Rep. 2016. PMID: 26948040 Free PMC article.
-
Viral proteases: Structure, mechanism and inhibition.Enzymes. 2021;50:301-333. doi: 10.1016/bs.enz.2021.09.004. Epub 2021 Nov 17. Enzymes. 2021. PMID: 34861941 Free PMC article.
References
-
- Allaire M., Chernaia M. M., Malcolm B. A., James M. N. 1994. Picornaviral 3C cysteine proteinases have a fold similar to chymotrypsin-like serine proteinases. Nature 369:72–76 - PubMed
-
- Bergmann E. M., et al. 1999. Crystal structure of an inhibitor complex of the 3C proteinase from hepatitis A virus (HAV) and implications for the polyprotein processing in HAV. Virology 265:153–163 - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources

