Stepwise unfolding of a β barrel protein by the AAA+ ClpXP protease
- PMID: 21821046
- PMCID: PMC3184388
- DOI: 10.1016/j.jmb.2011.07.041
Stepwise unfolding of a β barrel protein by the AAA+ ClpXP protease
Erratum in
- J Mol Biol. 2013 Apr 12;425(7):1241-3
Abstract
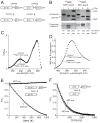
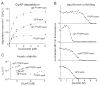
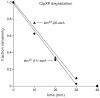
In the AAA+ ClpXP protease, ClpX uses the energy of ATP binding and hydrolysis to unfold proteins before translocating them into ClpP for degradation. For proteins with C-terminal ssrA tags, ClpXP pulls on the tag to initiate unfolding and subsequent degradation. Here, we demonstrate that an initial step in ClpXP unfolding of the 11-stranded β barrel of superfolder GFP-ssrA involves extraction of the C-terminal β strand. The resulting 10-stranded intermediate is populated at low ATP concentrations, which stall ClpXP unfolding, and at high ATP concentrations, which support robust degradation. To determine if stable unfolding intermediates cause low-ATP stalling, we designed and characterized circularly permuted GFP variants. Notably, stalling was observed for a variant that formed a stable 10-stranded intermediate but not for one in which this intermediate was unstable. A stepwise degradation model in which the rates of terminal-strand extraction, strand refolding or recapture, and unfolding of the 10-stranded intermediate all depend on the rate of ATP hydrolysis by ClpXP accounts for the observed changes in degradation kinetics over a broad range of ATP concentrations. Our results suggest that the presence or absence of unfolding intermediates will play important roles in determining whether forced enzymatic unfolding requires a minimum rate of ATP hydrolysis.
Copyright © 2011 Elsevier Ltd. All rights reserved.
Figures



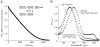
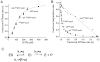
Similar articles
-
Protein unfolding by a AAA+ protease is dependent on ATP-hydrolysis rates and substrate energy landscapes.Nat Struct Mol Biol. 2008 Feb;15(2):139-45. doi: 10.1038/nsmb.1380. Epub 2008 Jan 27. Nat Struct Mol Biol. 2008. PMID: 18223658
-
Multistep substrate binding and engagement by the AAA+ ClpXP protease.Proc Natl Acad Sci U S A. 2020 Nov 10;117(45):28005-28013. doi: 10.1073/pnas.2010804117. Epub 2020 Oct 26. Proc Natl Acad Sci U S A. 2020. PMID: 33106413 Free PMC article.
-
Single-molecule denaturation and degradation of proteins by the AAA+ ClpXP protease.Proc Natl Acad Sci U S A. 2009 Nov 17;106(46):19340-5. doi: 10.1073/pnas.0910484106. Epub 2009 Nov 5. Proc Natl Acad Sci U S A. 2009. PMID: 19892734 Free PMC article.
-
ClpXP, an ATP-powered unfolding and protein-degradation machine.Biochim Biophys Acta. 2012 Jan;1823(1):15-28. doi: 10.1016/j.bbamcr.2011.06.007. Epub 2011 Jun 27. Biochim Biophys Acta. 2012. PMID: 21736903 Free PMC article. Review.
-
Structure and function of ClpXP, a AAA+ proteolytic machine powered by probabilistic ATP hydrolysis.Crit Rev Biochem Mol Biol. 2022 Apr;57(2):188-204. doi: 10.1080/10409238.2021.1979461. Epub 2021 Dec 19. Crit Rev Biochem Mol Biol. 2022. PMID: 34923891 Free PMC article. Review.
Cited by
-
Hysteresis as a Marker for Complex, Overlapping Landscapes in Proteins.J Phys Chem Lett. 2013 Jan 3;4(1):180-188. doi: 10.1021/jz301893w. Epub 2012 Dec 18. J Phys Chem Lett. 2013. PMID: 23525263 Free PMC article.
-
Engineering fluorescent protein substrates for the AAA+ Lon protease.Protein Eng Des Sel. 2013 Apr;26(4):299-305. doi: 10.1093/protein/gzs105. Epub 2013 Jan 28. Protein Eng Des Sel. 2013. PMID: 23359718 Free PMC article.
-
Structural basis of ClpXP recognition and unfolding of ssrA-tagged substrates.Elife. 2020 Oct 22;9:e61496. doi: 10.7554/eLife.61496. Elife. 2020. PMID: 33089779 Free PMC article.
-
The proteasome 19S cap and its ubiquitin receptors provide a versatile recognition platform for substrates.Nat Commun. 2020 Jan 24;11(1):477. doi: 10.1038/s41467-019-13906-8. Nat Commun. 2020. PMID: 31980598 Free PMC article.
-
Incomplete proteasomal degradation of green fluorescent proteins in the context of tandem fluorescent protein timers.Mol Biol Cell. 2016 Jan 15;27(2):360-70. doi: 10.1091/mbc.E15-07-0525. Epub 2015 Nov 25. Mol Biol Cell. 2016. PMID: 26609072 Free PMC article.
References
-
- Palombella VJ, Rando OJ, Goldberg AL, Maniatis T. The ubiquitin-proteasome pathway is required for processing the NF-κB1 precursor protein and the activation of NF-κB. Cell. 1994;78:773–785. - PubMed
-
- Lee C, Schwartz MP, Prakash S, Iwakura M, Matouschek A. ATP-dependent proteases degrade their substrates by processively unraveling them from the degradation signal. Mol Cell. 2001;7:627–637. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

