Cryptococcal titan cell formation is regulated by G-protein signaling in response to multiple stimuli
- PMID: 21821718
- PMCID: PMC3187071
- DOI: 10.1128/EC.05179-11
Cryptococcal titan cell formation is regulated by G-protein signaling in response to multiple stimuli
Abstract
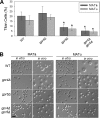
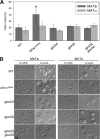
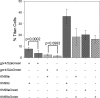
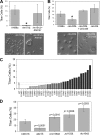
The titan cell is a recently described morphological form of the pathogenic fungus Cryptococcus neoformans. Occurring during the earliest stages of lung infection, titan cells are 5 to 10 times larger than the normal yeast-like cells, thereby resisting engulfment by lung phagocytes and favoring the persistence of infection. These enlarged cells exhibit an altered capsule structure, a thickened cell wall, increased ploidy, and resistance to nitrosative and oxidative stresses. We demonstrate that two G-protein-coupled receptors are important for induction of the titan cell phenotype: the Ste3a pheromone receptor (in mating type a cells) and the Gpr5 protein. Both receptors control titan cell formation through elements of the cyclic AMP (cAMP)/protein kinase A (PKA) pathway. This conserved signaling pathway, in turn, mediates its effect on titan cells through the PKA-regulated Rim101 transcription factor. Additional downstream effectors required for titan cell formation include the G(1) cyclin Pcl103, the Rho104 GTPase, and two GTPase-activating proteins, Gap1 and Cnc1560. These observations support developing models in which the PKA signaling pathway coordinately regulates many virulence-associated phenotypes in diverse human pathogens.
Figures

Similar articles
-
The Cryptococcus neoformans Titan cell is an inducible and regulated morphotype underlying pathogenesis.PLoS Pathog. 2018 May 18;14(5):e1006978. doi: 10.1371/journal.ppat.1006978. eCollection 2018 May. PLoS Pathog. 2018. PMID: 29775474 Free PMC article.
-
The Cryptococcus neoformans Rim101 transcription factor directly regulates genes required for adaptation to the host.Mol Cell Biol. 2014 Feb;34(4):673-84. doi: 10.1128/MCB.01359-13. Epub 2013 Dec 9. Mol Cell Biol. 2014. PMID: 24324006 Free PMC article.
-
The cAMP/Protein Kinase a Pathway Regulates Virulence and Adaptation to Host Conditions in Cryptococcus neoformans.Front Cell Infect Microbiol. 2019 Jun 18;9:212. doi: 10.3389/fcimb.2019.00212. eCollection 2019. Front Cell Infect Microbiol. 2019. PMID: 31275865 Free PMC article. Review.
-
The Cyclin Cln1 Controls Polyploid Titan Cell Formation following a Stress-Induced G2 Arrest in Cryptococcus.mBio. 2021 Oct 26;12(5):e0250921. doi: 10.1128/mBio.02509-21. Epub 2021 Oct 12. mBio. 2021. PMID: 34634930 Free PMC article.
-
Cryptococcal Titan Cells: When Yeast Cells Are All Grown up.Curr Top Microbiol Immunol. 2019;422:101-120. doi: 10.1007/82_2018_145. Curr Top Microbiol Immunol. 2019. PMID: 30406867 Review.
Cited by
-
How Environmental Fungi Cause a Range of Clinical Outcomes in Susceptible Hosts.J Mol Biol. 2019 Jul 26;431(16):2982-3009. doi: 10.1016/j.jmb.2019.05.003. Epub 2019 May 9. J Mol Biol. 2019. PMID: 31078554 Free PMC article. Review.
-
Morphology Changes in Human Fungal Pathogens upon Interaction with the Host.J Fungi (Basel). 2017;3(4):66. doi: 10.3390/jof3040066. Epub 2017 Dec 1. J Fungi (Basel). 2017. PMID: 29333431 Free PMC article.
-
cis- and trans-acting localization determinants of pH response regulator Rim13 in Saccharomyces cerevisiae.Eukaryot Cell. 2012 Oct;11(10):1201-9. doi: 10.1128/EC.00158-12. Epub 2012 Aug 3. Eukaryot Cell. 2012. PMID: 22865500 Free PMC article.
-
Fungal Strategies to Evade the Host Immune Recognition.J Fungi (Basel). 2017 Sep 23;3(4):51. doi: 10.3390/jof3040051. J Fungi (Basel). 2017. PMID: 29371567 Free PMC article. Review.
-
Shift and adapt: the costs and benefits of karyotype variations.Curr Opin Microbiol. 2015 Aug;26:130-6. doi: 10.1016/j.mib.2015.06.010. Epub 2015 Aug 25. Curr Opin Microbiol. 2015. PMID: 26321163 Free PMC article. Review.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

