Translation of HLA-HIV associations to the cellular level: HIV adapts to inflate CD8 T cell responses against Nef and HLA-adapted variant epitopes
- PMID: 21821798
- PMCID: PMC3183574
- DOI: 10.4049/jimmunol.1100691
Translation of HLA-HIV associations to the cellular level: HIV adapts to inflate CD8 T cell responses against Nef and HLA-adapted variant epitopes
Abstract
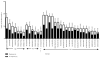
Strong statistical associations between polymorphisms in HIV-1 population sequences and carriage of HLA class I alleles have been widely used to identify possible sites of CD8 T cell immune selection in vivo. However, there have been few attempts to prospectively and systematically test these genetic hypotheses arising from population-based studies at a cellular, functional level. We assayed CD8 T cell epitope-specific IFN-γ responses in 290 individuals from the same cohort, which gave rise to 874 HLA-HIV associations in genetic analyses, taking into account autologous viral sequences and individual HLA genotypes. We found immunological evidence for 58% of 374 associations tested as sites of primary immune selection and identified up to 50 novel HIV-1 epitopes using this reverse-genomics approach. Many HLA-adapted epitopes elicited equivalent or higher-magnitude IFN-γ responses than did the nonadapted epitopes, particularly in Nef. At a population level, inclusion of all of the immunoreactive variant CD8 T cell epitopes in Gag, Pol, Nef, and Env suggested that HIV adaptation leads to an inflation of Nef-directed immune responses relative to other proteins. We concluded that HLA-HIV associations mark viral epitopes subject to CD8 T cell selection. These results can be used to guide functional studies of specific epitopes and escape mutations, as well as to test, train, and evaluate analytical models of viral escape and fitness. The inflation of Nef and HLA-adapted variant responses may have negative effects on natural and vaccine immunity against HIV and, therefore, has implications for diversity coverage approaches in HIV vaccine design.
Figures




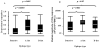


References
-
- Fischer W, Perkins S, Theiler J, Bhattacharya T, Yusim K, Funkhouser R, Kuiken C, Haynes B, Letvin NL, Walker BD, Hahn BH, Korber BT. Polyvalent vaccines for optimal coverage of potential T-cell epitopes in global HIV-1 variants. Nat Med. 2007;13:100–106. - PubMed
-
- Phillips RE, Rowland-Jones S, Nixon DF, Gotch FM, Edwards JP, Ogunlesi AO, Elvin JG, Rothbard JA, Bangham CR, Rizza CR, et al. Human immunodeficiency virus genetic variation that can escape cytotoxic T cell recognition. Nature. 1991;354:453–459. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
- AI069419/AI/NIAID NIH HHS/United States
- UM1 AI069424/AI/NIAID NIH HHS/United States
- AI27661/AI/NIAID NIH HHS/United States
- AI069450/AI/NIAID NIH HHS/United States
- U01 AI069477/AI/NIAID NIH HHS/United States
- U01 AI069502/AI/NIAID NIH HHS/United States
- U01 AI069513/AI/NIAID NIH HHS/United States
- AI68634/AI/NIAID NIH HHS/United States
- AI69467/AI/NIAID NIH HHS/United States
- UM1 AI069423/AI/NIAID NIH HHS/United States
- UM1 AI069494/AI/NIAID NIH HHS/United States
- AI68636/AI/NIAID NIH HHS/United States
- U01 AI069474/AI/NIAID NIH HHS/United States
- AI32782/AI/NIAID NIH HHS/United States
- U01 AI069434/AI/NIAID NIH HHS/United States
- AI069556/AI/NIAID NIH HHS/United States
- U01 AI069447/AI/NIAID NIH HHS/United States
- UM1 AI069501/AI/NIAID NIH HHS/United States
- UM1 AI069472/AI/NIAID NIH HHS/United States
- U01 AI069467/AI/NIAID NIH HHS/United States
- U01 AI069423/AI/NIAID NIH HHS/United States
- UM1 AI069513/AI/NIAID NIH HHS/United States
- AI69494/AI/NIAID NIH HHS/United States
- AI46370/AI/NIAID NIH HHS/United States
- U01 AI027661/AI/NIAID NIH HHS/United States
- U01 AI069465/AI/NIAID NIH HHS/United States
- UM1 AI069434/AI/NIAID NIH HHS/United States
- K24 AI064086/AI/NIAID NIH HHS/United States
- UM1 AI069411/AI/NIAID NIH HHS/United States
- UM1 AI069432/AI/NIAID NIH HHS/United States
- AI069501/AI/NIAID NIH HHS/United States
- AI069532/AI/NIAID NIH HHS/United States
- AI069474/AI/NIAID NIH HHS/United States
- AI069434/AI/NIAID NIH HHS/United States
- UM1 AI069495/AI/NIAID NIH HHS/United States
- AI38858/AI/NIAID NIH HHS/United States
- U01 AI069484/AI/NIAID NIH HHS/United States
- AI69423/AI/NIAID NIH HHS/United States
- UM1 AI069471/AI/NIAID NIH HHS/United States
- U01 AI069439/AI/NIAID NIH HHS/United States
- U01 AI069556/AI/NIAID NIH HHS/United States
- AI069513/AI/NIAID NIH HHS/United States
- AI069502/AI/NIAID NIH HHS/United States
- U01 AI069532/AI/NIAID NIH HHS/United States
- UM1 AI069439/AI/NIAID NIH HHS/United States
- AI069439/AI/NIAID NIH HHS/United States
- UM1 AI069484/AI/NIAID NIH HHS/United States
- UM1 AI068634/AI/NIAID NIH HHS/United States
- U01 AI069501/AI/NIAID NIH HHS/United States
- AI69411/AI/NIAID NIH HHS/United States
- AI34853/AI/NIAID NIH HHS/United States
- U01 AI046376/AI/NIAID NIH HHS/United States
- AI069452/AI/NIAID NIH HHS/United States
- AI069495/AI/NIAID NIH HHS/United States
- UM1 AI069477/AI/NIAID NIH HHS/United States
- AI-68636/AI/NIAID NIH HHS/United States
- U01 AI069432/AI/NIAID NIH HHS/United States
- AI069484/AI/NIAID NIH HHS/United States
- U01 AI069450/AI/NIAID NIH HHS/United States
- U01 AI038858/AI/NIAID NIH HHS/United States
- RR024975/RR/NCRR NIH HHS/United States
- U01 AI046370/AI/NIAID NIH HHS/United States
- AI046376/AI/NIAID NIH HHS/United States
- U01 AI068636/AI/NIAID NIH HHS/United States
- UM1 AI069474/AI/NIAID NIH HHS/United States
- AI69465/AI/NIAID NIH HHS/United States
- U01 AI034853/AI/NIAID NIH HHS/United States
- UM1 AI069452/AI/NIAID NIH HHS/United States
- AI069424/AI/NIAID NIH HHS/United States
- R01 AI060460/AI/NIAID NIH HHS/United States
- U01 AI069495/AI/NIAID NIH HHS/United States
- UM1 AI069556/AI/NIAID NIH HHS/United States
- P30 AI110527/AI/NIAID NIH HHS/United States
- U01 AI027673/AI/NIAID NIH HHS/United States
- AI64086/AI/NIAID NIH HHS/United States
- UM1 AI069502/AI/NIAID NIH HHS/United States
- U01 AI025859/AI/NIAID NIH HHS/United States
- UM1 AI069450/AI/NIAID NIH HHS/United States
- UM1 AI069532/AI/NIAID NIH HHS/United States
- AI27673/AI/NIAID NIH HHS/United States
- UM1 AI069465/AI/NIAID NIH HHS/United States
- U01 AI069424/AI/NIAID NIH HHS/United States
- U01 AI032782/AI/NIAID NIH HHS/United States
- UL1 RR024975/RR/NCRR NIH HHS/United States
- U01 AI069411/AI/NIAID NIH HHS/United States
- AI069472/AI/NIAID NIH HHS/United States
- UM1 AI069419/AI/NIAID NIH HHS/United States
- AI069432/AI/NIAID NIH HHS/United States
- AI069447/AI/NIAID NIH HHS/United States
- U01 AI069419/AI/NIAID NIH HHS/United States
- AI25859/AI/NIAID NIH HHS/United States
- U01 AI069494/AI/NIAID NIH HHS/United States
- UM1 AI069447/AI/NIAID NIH HHS/United States
- U01 AI069452/AI/NIAID NIH HHS/United States
- UM1 AI069467/AI/NIAID NIH HHS/United States
- AI069477/AI/NIAID NIH HHS/United States
- AI069471/AI/NIAID NIH HHS/United States
- U01 AI069471/AI/NIAID NIH HHS/United States
- U01 AI069472/AI/NIAID NIH HHS/United States
- U01 AI068634/AI/NIAID NIH HHS/United States
- UM1 AI068636/AI/NIAID NIH HHS/United States
LinkOut - more resources
Full Text Sources
Medical
Research Materials

