The genome of the extremophile crucifer Thellungiella parvula
- PMID: 21822265
- PMCID: PMC3586812
- DOI: 10.1038/ng.889
The genome of the extremophile crucifer Thellungiella parvula
Abstract
Thellungiella parvula is related to Arabidopsis thaliana and is endemic to saline, resource-poor habitats, making it a model for the evolution of plant adaptation to extreme environments. Here we present the draft genome for this extremophile species. Exclusively by next generation sequencing, we obtained the de novo assembled genome in 1,496 gap-free contigs, closely approximating the estimated genome size of 140 Mb. We anchored these contigs to seven pseudo chromosomes without the use of maps. We show that short reads can be assembled to a near-complete chromosome level for a eukaryotic species lacking prior genetic information. The sequence identifies a number of tandem duplications that, by the nature of the duplicated genes, suggest a possible basis for T. parvula's extremophile lifestyle. Our results provide essential background for developing genomically influenced testable hypotheses for the evolution of environmental stress tolerance.
Conflict of interest statement
The authors declare no competing financial interests.
Figures



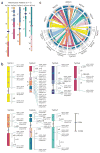
References
-
- Al-Shehbaz IA, O’Kane SL. Placement of Arabidopsis parvula in Thellungiella (Brassicaceae) Novon. 1995;5:309–310.
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

