A single methyltransferase YefA (RlmCD) catalyses both m5U747 and m5U1939 modifications in Bacillus subtilis 23S rRNA
- PMID: 21824914
- PMCID: PMC3241648
- DOI: 10.1093/nar/gkr626
A single methyltransferase YefA (RlmCD) catalyses both m5U747 and m5U1939 modifications in Bacillus subtilis 23S rRNA
Abstract
Methyltransferases that use S-adenosylmethionine (AdoMet) as a cofactor to catalyse 5-methyl uridine (m(5)U) formation in tRNAs and rRNAs are widespread in Bacteria and Eukaryota, and are also found in certain Archaea. These enzymes belong to the COG2265 cluster, and the Gram-negative bacterium Escherichia coli possesses three paralogues. These comprise the methyltransferases TrmA that targets U54 in tRNAs, RlmC that modifies U747 in 23S rRNA and RlmD that is specific for U1939 in 23S rRNA. The tRNAs and rRNAs of the Gram-positive bacterium Bacillus subtilis have the same three m(5)U modifications. However, as previously shown, the m(5)U54 modification in B. subtilis tRNAs is catalysed in a fundamentally different manner by the folate-dependent enzyme TrmFO, which is unrelated to the E. coli TrmA. Here, we show that methylation of U747 and U1939 in B. subtilis rRNA is catalysed by a single enzyme, YefA that is a COG2265 member. A recombinant version of YefA functions in an E. coli m(5)U-null mutant adding the same two rRNA methylations. The findings suggest that during evolution, COG2265 enzymes have undergone a series of changes in target specificity and that YefA is closer to an archetypical m(5)U methyltransferase. To reflect its dual specificity, YefA is renamed RlmCD.
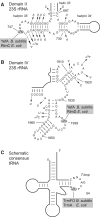
Figures


Similar articles
-
Specificity shifts in the rRNA and tRNA nucleotide targets of archaeal and bacterial m5U methyltransferases.RNA. 2011 Jan;17(1):45-53. doi: 10.1261/rna.2323411. Epub 2010 Nov 4. RNA. 2011. PMID: 21051506 Free PMC article.
-
Identifying the methyltransferases for m(5)U747 and m(5)U1939 in 23S rRNA using MALDI mass spectrometry.Nucleic Acids Res. 2003 Aug 15;31(16):4738-46. doi: 10.1093/nar/gkg657. Nucleic Acids Res. 2003. PMID: 12907714 Free PMC article.
-
The flavoprotein Mcap0476 (RlmFO) catalyzes m5U1939 modification in Mycoplasma capricolum 23S rRNA.Nucleic Acids Res. 2014 Jul;42(12):8073-82. doi: 10.1093/nar/gku518. Epub 2014 Jun 17. Nucleic Acids Res. 2014. PMID: 24939895 Free PMC article.
-
RlmCD-mediated U747 methylation promotes efficient G748 methylation by methyltransferase RlmAII in 23S rRNA in Streptococcus pneumoniae; interplay between two rRNA methylations responsible for telithromycin susceptibility.Nucleic Acids Res. 2015 Oct 15;43(18):8964-72. doi: 10.1093/nar/gkv609. Epub 2015 Sep 13. Nucleic Acids Res. 2015. PMID: 26365244 Free PMC article.
-
Ribosomal RNA guanine-(N2)-methyltransferases and their targets.Nucleic Acids Res. 2007;35(7):2295-301. doi: 10.1093/nar/gkm104. Epub 2007 Mar 27. Nucleic Acids Res. 2007. PMID: 17389639 Free PMC article. Review.
Cited by
-
Methylation of 23S rRNA nucleotide G748 by RlmAII methyltransferase renders Streptococcus pneumoniae telithromycin susceptible.Antimicrob Agents Chemother. 2013 Aug;57(8):3789-96. doi: 10.1128/AAC.00164-13. Epub 2013 May 28. Antimicrob Agents Chemother. 2013. PMID: 23716046 Free PMC article.
-
Complete list of canonical post-transcriptional modifications in the Bacillus subtilis ribosome and their link to RbgA driven large subunit assembly.Nucleic Acids Res. 2024 Oct 14;52(18):11203-11217. doi: 10.1093/nar/gkae626. Nucleic Acids Res. 2024. PMID: 39036956 Free PMC article.
-
Mapping of ribosomal 23S ribosomal RNA modifications in Clostridium sporogenes.RNA Biol. 2018;15(8):1060-1070. doi: 10.1080/15476286.2018.1486662. Epub 2018 Aug 13. RNA Biol. 2018. PMID: 29947286 Free PMC article.
-
Complete list of canonical post-transcriptional modifications in the Bacillus subtilis ribosome and their link to RbgA driven large subunit assembly.bioRxiv [Preprint]. 2024 May 11:2024.05.10.593627. doi: 10.1101/2024.05.10.593627. bioRxiv. 2024. Update in: Nucleic Acids Res. 2024 Oct 14;52(18):11203-11217. doi: 10.1093/nar/gkae626. PMID: 38765983 Free PMC article. Updated. Preprint.
-
RNA-methyltransferase TrmA is a dual-specific enzyme responsible for C5-methylation of uridine in both tmRNA and tRNA.RNA Biol. 2013 Apr;10(4):572-8. doi: 10.4161/rna.24327. Epub 2013 Apr 1. RNA Biol. 2013. PMID: 23603891 Free PMC article.
References
-
- Björk GR, Hagervall TG. Transfer RNA modification. In: Böck A, Curtis R, Kaper JB, Neidhardt T, Nyström T, Squires C, editors. Escherichia coli and Salmonella. Washington DC: ASM Press; 2005. p. 4.6.2.
-
- Grosjean H. DNA and RNA Modification Enzymes: Structure, Mechanism, Function and Evolution. Austin Texas: Landes Biosciences; 2009.
-
- Urbonavičius J, Auxilien S, Walbott H, Trachana K, Golinelli-Pimpaneau B, Brochier-Armanet C, Grosjean H. Acquisition of a bacterial RumA-type tRNA(uracil-54, C5)-methyltransferase by Archaea through an ancient horizontal gene transfer. Mol. Microbiol. 2008;67:323–335. - PubMed
-
- Ofengand J, Del Campo M. Modified nucleotides of E. coli ribosomal RNA. In: Böck A, Curtis R, Kaper JB, Neidhardt T, Nyström T, Squires C, editors. Escherichia coli and Salmonella. Washington DC: ASM Press; 2005. p. 4.6.1.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

