Genome-wide methylation analysis identifies genes specific to breast cancer hormone receptor status and risk of recurrence
- PMID: 21825015
- PMCID: PMC3308629
- DOI: 10.1158/0008-5472.CAN-11-1630
Genome-wide methylation analysis identifies genes specific to breast cancer hormone receptor status and risk of recurrence
Abstract
To better understand the biology of hormone receptor-positive and-negative breast cancer and to identify methylated gene markers of disease progression, we carried out a genome-wide methylation array analysis on 103 primary invasive breast cancers and 21 normal breast samples, using the Illumina Infinium HumanMethylation27 array that queried 27,578 CpG loci. Estrogen and/or progesterone receptor-positive tumors displayed more hypermethylated loci than estrogen receptor (ER)-negative tumors. However, the hypermethylated loci in ER-negative tumors were clustered closer to the transcriptional start site compared with ER-positive tumors. An ER-classifier set of CpG loci was identified, which independently partitioned primary tumors into ER subtypes. A total of 40 (32 novel and 8 previously known) CpG loci showed differential methylation specific to either ER-positive or ER-negative tumors. Each of the 40 ER subtype-specific loci was validated in silico, using an independent, publicly available methylome dataset from the Cancer Genome Atlas. In addition, we identified 100 methylated CpG loci that were significantly associated with disease progression; the majority of these loci were informative particularly in ER-negative breast cancer. Overall, the set was highly enriched in homeobox containing genes. This pilot study shows the robustness of the breast cancer methylome and illustrates its potential to stratify and reveal biological differences between ER subtypes of breast cancer. Furthermore, it defines candidate ER-specific markers and identifies potential markers predictive of outcome within ER subgroups.
Conflict of interest statement
No conflict of interest.
Figures

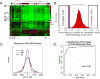



References
-
- Carlson RW, Allred DC, Anderson BO, Burstein HJ, Carter WB, Edge SB, et al. Breast cancer. Clinical practice guidelines in oncology. J Natl Compr Canc Netw. 2009;7:122–92. - PubMed
-
- Hammond ME, Hayes DF, Dowsett M, Allred DC, Hagerty KL, Badve S, et al. American Society of Clinical Oncology/College Of American Pathologists guideline recommendations for immunohistochemical testing of estrogen and progesterone receptors in breast cancer. J Clin Oncol. 2010;28:2784–95. - PMC - PubMed
-
- Harris L, Fritsche H, Mennel R, Norton L, Ravdin P, Taube S, et al. American Society of Clinical Oncology 2007 update of recommendations for the use of tumor markers in breast cancer. J Clin Oncol. 2007;25:5287–312. - PubMed
-
- Wolff AC, Hammond ME, Schwartz JN, Hagerty KL, Allred DC, Cote RJ, et al. American Society of Clinical Oncology/College of American Pathologists guideline recommendations for human epidermal growth factor receptor 2 testing in breast cancer. J Clin Oncol. 2007;25:118–45. - PubMed
-
- Fisher B, Dignam J, Tan-Chiu E, Anderson S, Fisher ER, Wittliff JL, et al. Prognosis and treatment of patients with breast tumors of one centimeter or less and negative axillary lymph nodes. J Natl Cancer Inst. 2001;93:112–20. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials

