Characterization of STAT6 target genes in human B cells and lung epithelial cells
- PMID: 21828071
- PMCID: PMC3190958
- DOI: 10.1093/dnares/dsr025
Characterization of STAT6 target genes in human B cells and lung epithelial cells
Abstract
Using ChIP Seq, we identified 556 and 467 putative STAT6 target sites in the Burkitt's lymphoma cell line Ramos and in the normal lung epithelial cell line BEAS2B, respectively. We also examined the positions and expression of transcriptional start sites (TSSs) in these cells using our TSS Seq method. We observed that 44 and 132 genes in Ramos and BEAS2B, respectively, had STAT6 binding sites in proximal regions of their previously reported TSSs that were up-regulated at the transcriptional level. In addition, 406 and 109 of the STAT6 target sites in Ramos and BEAS2B, respectively, were located in proximal regions of previously uncharacterized TSSs. The target genes identified in Ramos and BEAS2B cells in this study and in Th2 cells in previous studies rarely overlapped and differed in their identity. Interestingly, ChIP Seq analyses of histone modifications and RNA polymerase II revealed that chromatin formed an active structure in regions surrounding the STAT6 binding sites; this event also frequently occurred in different cell types, although neither STAT6 binding nor TSS induction was observed. The rough landscape of STAT6-responsive sites was found to be shaped by chromatin structure, but distinct cellular responses were mainly mediated by distinct sets of transcription factors.
Figures



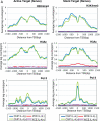


References
-
- Kaplan M.H., Schindler U., Smiley S.T., Grusby M.J. Stat6 is required for mediating responses to IL-4 and for development of Th2 cells. Immunity. 1996;4:313–19. - PubMed
-
- Shirakawa I., Deichmann K.A., Izuhara I., Mao I., Adra C.N., Hopkin J.M. Atopy and asthma: genetic variants of IL-4 and IL-13 signalling. Immunol. Today. 2000;21:60–4. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

