Definition of membrane topology and identification of residues important for transport in subunit a of the vacuolar ATPase
- PMID: 21832060
- PMCID: PMC3186431
- DOI: 10.1074/jbc.M111.273409
Definition of membrane topology and identification of residues important for transport in subunit a of the vacuolar ATPase
Abstract
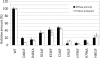
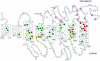
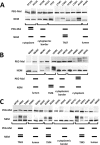
Subunit a of the vacuolar H(+)-ATPases plays an important role in proton transport. This membrane-integral 100-kDa subunit is thought to form or contribute to proton-conducting hemichannels that allow protons to gain access to and leave buried carboxyl groups on the proteolipid subunits (c, c', and c″) during proton translocation. We previously demonstrated that subunit a contains a large N-terminal cytoplasmic domain followed by a C-terminal domain containing eight transmembrane (TM) helices. TM7 contains a buried arginine residue (Arg-735) that is essential for proton transport and is located on a helical face that interacts with the proteolipid ring. To further define the topology of the C-terminal domain, the accessibility of 30 unique cysteine residues to the membrane-permeant reagent N-ethylmaleimide and the membrane-impermeant reagent polyethyleneglycol maleimide was determined. The results further define the borders of transmembrane segments in subunit a. To identify additional buried polar and charged residues important in proton transport, 25 sites were individually mutated to hydrophobic amino acids, and the effect on proton transport was determined. These and previous results identify a set of residues important for proton transport located on the cytoplasmic half of TM7 and TM8 and the lumenal half of TM3, TM4, and TM7. Based upon these data, we propose a tentative model in which the cytoplasmic hemichannel is located at the interface of TM7 and TM8 of subunit a and the proteolipid ring, whereas the lumenal hemichannel is located within subunit a at the interface of TM3, TM4, and TM7.
Figures



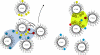
Similar articles
-
TM2 but not TM4 of subunit c'' interacts with TM7 of subunit a of the yeast V-ATPase as defined by disulfide-mediated cross-linking.J Biol Chem. 2004 Oct 22;279(43):44628-38. doi: 10.1074/jbc.M407345200. Epub 2004 Aug 18. J Biol Chem. 2004. PMID: 15322078
-
Analysis of the membrane topology of transmembrane segments in the C-terminal hydrophobic domain of the yeast vacuolar ATPase subunit a (Vph1p) by chemical modification.J Biol Chem. 2008 Jul 25;283(30):20696-702. doi: 10.1074/jbc.M803258200. Epub 2008 May 28. J Biol Chem. 2008. PMID: 18508769 Free PMC article.
-
Interacting helical surfaces of the transmembrane segments of subunits a and c' of the yeast V-ATPase defined by disulfide-mediated cross-linking.J Biol Chem. 2003 Oct 24;278(43):41908-13. doi: 10.1074/jbc.M308026200. Epub 2003 Aug 12. J Biol Chem. 2003. PMID: 12917411
-
Structural aspects of the gastric H,K ATPase.Ann N Y Acad Sci. 1997 Nov 3;834:65-76. doi: 10.1111/j.1749-6632.1997.tb52226.x. Ann N Y Acad Sci. 1997. PMID: 9405786 Review.
-
Subunit structure, function, and arrangement in the yeast and coated vesicle V-ATPases.J Bioenerg Biomembr. 2003 Aug;35(4):291-9. doi: 10.1023/a:1025720713747. J Bioenerg Biomembr. 2003. PMID: 14635775 Review.
Cited by
-
Structural and functional understanding of disease-associated mutations in V-ATPase subunit a1 and other isoforms.Front Mol Neurosci. 2023 Jul 3;16:1135015. doi: 10.3389/fnmol.2023.1135015. eCollection 2023. Front Mol Neurosci. 2023. PMID: 37465367 Free PMC article. Review.
-
Organelle acidification negatively regulates vacuole membrane fusion in vivo.Sci Rep. 2016 Jul 1;6:29045. doi: 10.1038/srep29045. Sci Rep. 2016. PMID: 27363625 Free PMC article.
-
The a subunit isoforms of vacuolar-type proton ATPase exhibit differential distribution in mouse perigastrulation embryos.Sci Rep. 2022 Aug 10;12(1):13590. doi: 10.1038/s41598-022-18002-4. Sci Rep. 2022. PMID: 35948619 Free PMC article.
-
Regulation of V-ATPase assembly and function of V-ATPases in tumor cell invasiveness.Biochim Biophys Acta. 2016 Aug;1857(8):1213-1218. doi: 10.1016/j.bbabio.2016.02.010. Epub 2016 Feb 22. Biochim Biophys Acta. 2016. PMID: 26906430 Free PMC article. Review.
-
Probing the proton channels in subunit N of Complex I from Escherichia coli through intra-subunit cross-linking.Biochim Biophys Acta. 2016 Dec;1857(12):1840-1848. doi: 10.1016/j.bbabio.2016.09.005. Epub 2016 Sep 12. Biochim Biophys Acta. 2016. PMID: 27632419 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

