Engineering the respiratory complex I to energy-converting NADPH:ubiquinone oxidoreductase
- PMID: 21832062
- PMCID: PMC3186356
- DOI: 10.1074/jbc.M111.274571
Engineering the respiratory complex I to energy-converting NADPH:ubiquinone oxidoreductase
Abstract
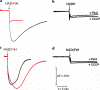
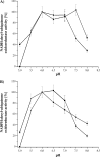
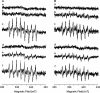
The respiratory complex I couples the electron transfer from NADH to ubiquinone with a translocation of protons across the membrane. Its nucleotide-binding site is made up of a unique Rossmann fold to accommodate the binding of the substrate NADH and of the primary electron acceptor flavin mononucleotide. Binding of NADH includes interactions of the hydroxyl groups of the adenosine ribose with a conserved glutamic acid residue. Structural analysis revealed that due to steric hindrance and electrostatic repulsion, this residue most likely prevents the binding of NADPH, which is a poor substrate of the complex. We produced several variants with mutations at this position exhibiting up to 200-fold enhanced catalytic efficiency with NADPH. The reaction of the variants with NAD(P)H is coupled with proton translocation in an inhibitor-sensitive manner. Thus, we have created an energy-converting NADPH:ubiquinone oxidoreductase, an activity so far not found in nature. Remarkably, the oxidation of NAD(P)H by the variants leads to an enhanced production of reactive oxygen species.
Figures


Similar articles
-
Spin labeling of the Escherichia coli NADH ubiquinone oxidoreductase (complex I).Biochim Biophys Acta. 2010 Dec;1797(12):1894-900. doi: 10.1016/j.bbabio.2010.10.013. Epub 2010 Oct 16. Biochim Biophys Acta. 2010. PMID: 20959113
-
Iron-sulfur cluster N2 of the Escherichia coli NADH:ubiquinone oxidoreductase (complex I) is located on subunit NuoB.J Biol Chem. 2003 Nov 28;278(48):47602-9. doi: 10.1074/jbc.M308967200. Epub 2003 Sep 15. J Biol Chem. 2003. PMID: 12975362
-
Energy conversion, redox catalysis and generation of reactive oxygen species by respiratory complex I.Biochim Biophys Acta. 2016 Jul;1857(7):872-83. doi: 10.1016/j.bbabio.2015.12.009. Epub 2015 Dec 22. Biochim Biophys Acta. 2016. PMID: 26721206 Free PMC article. Review.
-
On the mechanism of respiratory complex I.J Bioenerg Biomembr. 2014 Aug;46(4):255-68. doi: 10.1007/s10863-014-9566-8. Epub 2014 Jul 15. J Bioenerg Biomembr. 2014. PMID: 25022766 Review.
-
Nucleotide-induced conformational changes in the Escherichia coli NADH:ubiquinone oxidoreductase (complex I).Biochem Soc Trans. 2008 Oct;36(Pt 5):971-5. doi: 10.1042/BST0360971. Biochem Soc Trans. 2008. PMID: 18793172
Cited by
-
Changing the Electron Acceptor Specificity of Rhodobacter capsulatus Formate Dehydrogenase from NAD+ to NADP.Int J Mol Sci. 2023 Nov 8;24(22):16067. doi: 10.3390/ijms242216067. Int J Mol Sci. 2023. PMID: 38003259 Free PMC article.
-
Reduction of the off-pathway iron-sulphur cluster N1a of Escherichia coli respiratory complex I restrains NAD+ dissociation.Sci Rep. 2017 Aug 18;7(1):8754. doi: 10.1038/s41598-017-09345-4. Sci Rep. 2017. PMID: 28821859 Free PMC article.
-
Cofactor Specificity Engineering of Streptococcus mutans NADH Oxidase 2 for NAD(P)(+) Regeneration in Biocatalytic Oxidations.Comput Struct Biotechnol J. 2014 Feb 26;9:e201402005. doi: 10.5936/csbj.201402005. eCollection 2014. Comput Struct Biotechnol J. 2014. PMID: 24757503 Free PMC article.
-
A mechanism to prevent production of reactive oxygen species by Escherichia coli respiratory complex I.Nat Commun. 2019 Jun 11;10(1):2551. doi: 10.1038/s41467-019-10429-0. Nat Commun. 2019. PMID: 31186428 Free PMC article.
-
Arginase expression modulates nitric oxide production in Leishmania (Leishmania) amazonensis.PLoS One. 2017 Nov 14;12(11):e0187186. doi: 10.1371/journal.pone.0187186. eCollection 2017. PLoS One. 2017. PMID: 29135983 Free PMC article.
References
-
- Weiss H., Friedrich T., Hofhaus G., Preis D. (1991) Eur. J. Biochem. 197, 563–576 - PubMed
-
- Walker J. E. (1992) Q. Rev. Biophys. 25, 253–324 - PubMed
-
- Ohnishi T. (1998) Biochim. Biophys. Acta 1364, 186–206 - PubMed
-
- Friedrich T. (2001) J. Bioenerg. Biomembr. 33, 169–177 - PubMed
-
- Yagi T., Matsuno-Yagi A. (2003) Biochemistry 42, 2266–2274 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

