Polymorphic triple beta-sheet structures contribute to amide hydrogen/deuterium (H/D) exchange protection in the Alzheimer amyloid beta42 peptide
- PMID: 21832091
- PMCID: PMC3190809
- DOI: 10.1074/jbc.M111.241141
Polymorphic triple beta-sheet structures contribute to amide hydrogen/deuterium (H/D) exchange protection in the Alzheimer amyloid beta42 peptide
Abstract
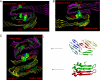
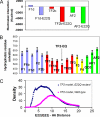
Characterization of the polymorphic structural range of Aβ oligomers is important to the understanding of the mechanisms of toxicity. Yet for highly polymorphic ensembles, experimental structural elucidation is difficult. Here, we use a combination of NMR solvent protection experiments and computational structural screening to identify major species in the amyloid conformational ensemble. We examined the polymorphic pentamer and fibril seeds of Aβ42 and its mutants and compared the theoretical backbone amide protection obtained from simulations with experimental hydrogen/deuterium (H/D) exchange protection ratio. We observed that highly flexible pentamers do not share structural similarities with fibril seed oligomers, except the turn regions. We found that a novel amyloid structural motif of a triple β-sheet, with the N-terminal residues interacting with the core (Lys(17)-Glu(22)) β-sheet region, correlates with H/D exchange protection. The triple β-sheet Aβ42 oligomer has a minimal exposure of hydrophobic residues and is further stabilized by the E22Q (Dutch) mutation in Alzheimer disease. The experimental H/D exchange solvent protection ratio implies that triple β-sheet fibrils and globulomers could coexist in the Aβ42 ensemble, pointing to a broad heterogeneous aggregate population. Our results suggest that an approach that combines computational modeling with NMR protection data can be a useful strategy for obtaining clues to the preferred conformational species of the assemblies in solution and help in alleviating experimental difficulties and consequently possible errors in the exchange data for Aβ42 fibrils.
Figures
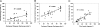
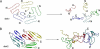




References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

